Cell Proliferation Tracer: Principles, Fluorescent Staining, and Dye Selection
Cell staining for proliferation analysis is not only about making cells visible. In many studies, researchers need to know whether cells are actually dividing, how many rounds of division have occurred, whether only part of a population is proliferating, and whether that behavior changes after stimulation, treatment, co-culture, or environmental shift. This is where a cell proliferation tracer becomes especially useful. Unlike a general fluorescent label that mainly reports location or short-term cell status, a proliferation tracer is designed to support division tracking over time. In practice, that means the fluorescent signal must be introduced in a way that is stable enough to remain associated with the labeled cell population, yet interpretable enough to reveal how that signal changes as cells divide.
Introduction to Cell Proliferation Tracers
A cell proliferation tracer is a fluorescent dyes or fluorescent probes used to label a cell population so that subsequent fluorescence changes can be used to assess cell division over time. Its value lies not simply in making cells fluorescent, but in turning fluorescence into an interpretable record of proliferative behavior. In typical workflows, cells are labeled before stimulation, treatment, or co-culture, and the signal is then analyzed after one or more rounds of division. As daughter cells are generated, the tracer signal changes in a way that helps researchers estimate generational progression, compare proliferative responses between samples, and identify whether the whole population or only a subset of cells is actively dividing. This makes a proliferation tracer a functional readout of division history rather than a simple structural stain. For users comparing available reagent options, cell proliferation tracer fluorescent probes can be reviewed according to assay design, signal behavior, and workflow compatibility.
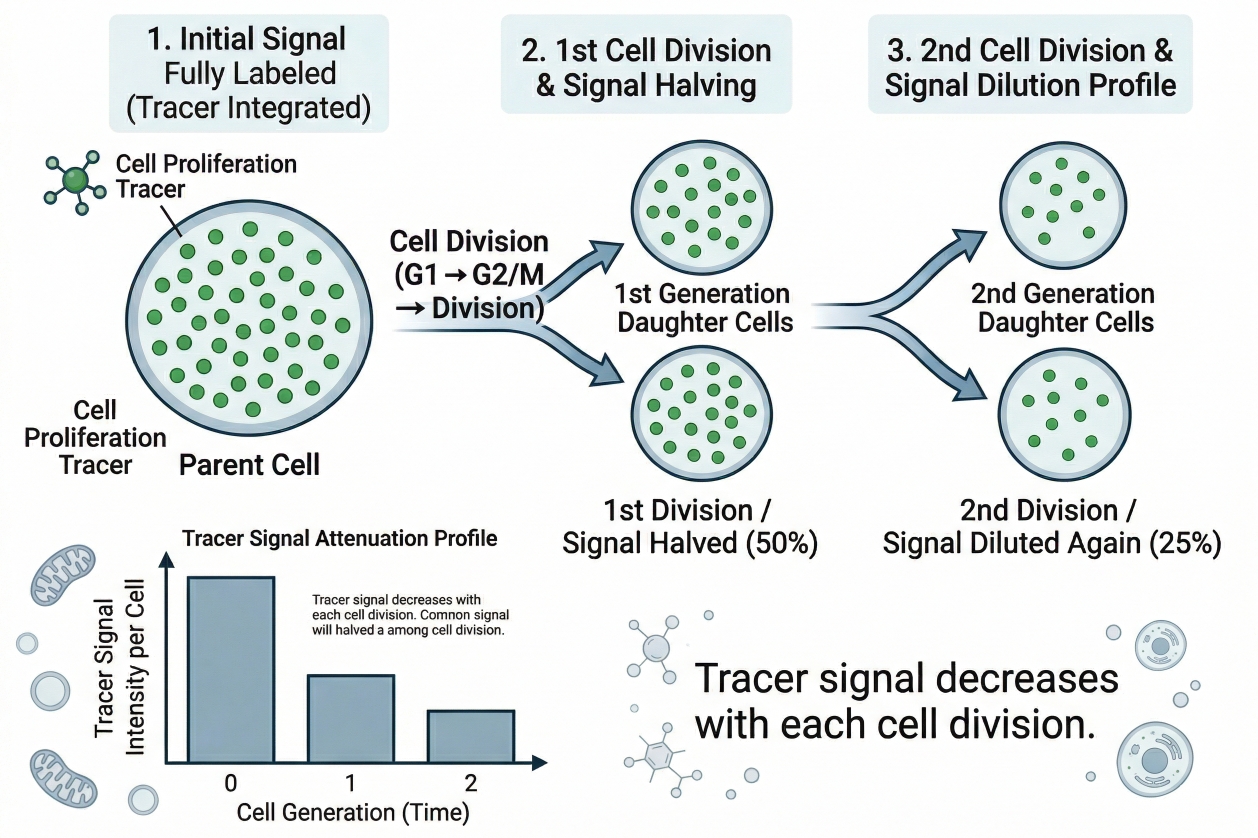 Fig. 1. Tracer signal decreases with each cell division (BOC Sciences Authorized).
Fig. 1. Tracer signal decreases with each cell division (BOC Sciences Authorized).
Cell proliferation tracers are therefore best understood as analytical tools rather than simple fluorescent labels. Their value depends on more than whether the initial staining looks bright. What matters is how the tracer signal is generated, how reliably it is retained, how clearly it can support multigeneration analysis, and whether decreasing fluorescence can be interpreted as true division-linked dilution rather than staining loss, handling-related variation, or other non-biological effects. They should also be considered within the broader context of cell and organelle stains, because proliferation experiments are often designed alongside other fluorescent workflows instead of being performed in isolation. In practical assay planning, tracer selection is rarely determined by color alone. It depends on how well the signal fits the analytical platform, the cell system, the expected assay duration, and the biological question the experiment is meant to answer.
Difference Between Proliferation Tracers and General Cell Stains
General fluorescent stains are often used to label location, viability-associated features, organelles, or short-term cellular state. A proliferation tracer serves a different purpose. Its value comes from stable labeling and interpretable signal change across divisions. That difference matters because a stain that is bright and easy to see is not automatically suitable for proliferation analysis. For proliferation tracing, the initial stain must be sufficiently uniform, sufficiently retained, and sufficiently compatible with ongoing cell behavior for the signal to remain analytically meaningful after cells begin dividing. This is why proliferation tracers should be evaluated as part of a full staining and readout strategy rather than chosen as if they were interchangeable with routine fluorescent cell labels.
Main Applications of Proliferation Tracing
Proliferation tracers are widely used when researchers need to examine division behavior in stimulated cell populations, compare proliferative responses between treatment groups, track heterogeneity in mixed populations, or connect proliferation status with phenotype markers in multicolor workflows. They are particularly helpful when a simple increase or decrease in total cell number is not enough to answer the scientific question. In those situations, the tracer can help distinguish whether the whole population is dividing modestly, whether only a subset is proliferating strongly, or whether one condition produces a broader spread of division histories than another. The technique is therefore valuable not only for measuring cell growth but for making cell population behavior more interpretable.
Need help choosing the right cell proliferation tracer?
We can help compare tracer chemistry, staining strategy, and readout design for flow cytometry and fluorescence-based proliferation analysis.
Basic Principles of Proliferation Tracer Fluorescent Staining
The usefulness of a proliferation tracer depends on how the fluorescent staining step is designed and how the resulting signal behaves after cells begin dividing. A well-designed tracer assay does not rely on fluorescence visibility alone. It relies on the relationship between initial labeling quality, intracellular retention, signal dilution, and the analytical platform used to read the result. Understanding these principles early helps prevent a common mistake in proliferation experiments: assuming that a fluorescently labeled cell population automatically produces interpretable proliferation data. In reality, the meaning of the tracer signal depends on how the stain was introduced, how consistently it was distributed, and how carefully downstream analysis is performed.
- Cell labeling and intracellular retention: A useful proliferation tracer must label cells in a way that is sufficiently stable for the fluorescent signal to remain associated with the cell population over the course of the assay. In practice, this means the staining approach should support retention during washing, recovery, culture, and repeated cell division. A signal that leaks rapidly or redistributes unpredictably can look fluorescent at the start of the experiment while still being a poor tracer for proliferation analysis.
- Signal dilution during cell division: In dye-dilution workflows, fluorescence becomes progressively reduced as cells divide. This reduction is valuable because it transforms the stain into a readout of division history rather than static labeling. The analytical goal is therefore not simply to observe a weaker signal over time, but to understand whether that change is consistent with division-linked dilution across the labeled population.
- Relationship between initial staining and data quality: The first staining step strongly influences the final analytical quality of the assay. If the starting signal is uneven, too broad, or unstable, later generations become difficult to resolve and interpret. A narrow and consistent parent population is usually much more valuable than a merely bright one, because resolution across later time points depends on the quality of the initial labeling event.
- Interpretation limits of fluorescent tracers: A proliferation tracer can support strong conclusions about division-linked signal behavior, but it does not automatically explain why a cell population divided, what signaling pathways were activated, or whether the observed differences are independent of cell health, survival, or selection effects. The tracer should therefore be treated as one readout within a broader experimental framework rather than a complete substitute for careful biological interpretation.
Common Types of Cell Proliferation Tracer Dyes and Probes
Cell proliferation tracers are best understood by staining route and use case rather than by name alone. Some fluorescent reagents are more naturally positioned as classic dye-dilution tools, while others are better thought of as general cell-tracking reagents that may support proliferation-related interpretation in selected workflows. The most practical way to choose among them is to ask what the assay needs the signal to do over time. Does it need to remain strongly associated with cells through multiple divisions, integrate into a multicolor panel, support live-cell handling, or provide a more general fluorescent history of labeled cells? Those questions are often more useful than starting with catalog familiarity alone.
Amine-Reactive Proliferation Dyes
Amine-reactive proliferation dyes are among the best-known routes for division tracking because they are designed to label cells in a way that supports stable intracellular retention and interpretable dilution across cell generations. In practice, these reagents are often selected when users want clear parent-population labeling, multigeneration readout, and compatibility with common proliferation analysis workflows. Their strength lies in the relationship between initial staining consistency and downstream generation resolution. These dyes are most useful when the assay is truly intended to follow division history rather than simply confirm that cells remain fluorescent over time.
Spectrally Distinct Fluorescent Tracers
Not all proliferation-compatible tracers occupy the same spectral space, and this becomes important as soon as the assay involves phenotype markers, reporter channels, viability readouts, or additional fluorescent stains. Spectrally distinct tracers give users flexibility when the green channel is already occupied or when a lower-conflict channel is needed for a multicolor panel. In the product context provided here, CytoTrace™ Orange CMTMR is a relevant example of a fluorescent cell tracer that may fit workflows requiring orange-channel allocation rather than conventional green-space usage. Spectral selection is therefore not just about appearance. It directly affects panel compatibility, compensation burden, and the interpretability of the final analysis.
Membrane-Associated and General Cell Tracking Dyes
Some fluorescent tracers are better thought of as broader cell-tracking tools rather than classic proliferation-only reagents. They may still contribute to proliferation-related workflows, especially when researchers need to follow labeled cells over time, distinguish a pre-labeled population from unlabeled cells, or combine tracking with imaging-based observation. Examples in this broader category can include compounds such as Green CMFDA, 5(6)-Carboxyfluorescein diacetate, 6-Carboxyfluorescein diacetate, and 6-Carboxyfluorescein diacetate succinimidyl ester, each of which highlights a different balance between intracellular staining behavior, signal persistence, and workflow role. These reagents can be useful, but they should not all be assumed to function identically as multigeneration proliferation tracers.
Fluorescent Probes for Division Tracking Workflows
In practical assay development, users often compare dyes, cell-tracking reagents, and fluorescent probes within the same planning stage because the final question is not simply what the reagent is called, but whether it produces a readable division-linked signal in the chosen system. Some reagents, such as 7-Amino-4-chloromethylcoumarin or Lucifer Yellow derivatives, may be more useful in selected tracer or loading workflows rather than as standard generational proliferation dyes. Others, such as Biocytin hydrazide or Lucifer yellow cadaverine, are more naturally associated with transport, loading, or tracking-related contexts. This is why selection should begin with assay purpose and readout logic, then move toward reagent class and spectral choice rather than the reverse.
Fluorescent Staining Strategies for Proliferation Analysis
Fluorescent staining design is one of the most decisive parts of any proliferation tracer experiment. Even a chemically suitable tracer can produce weak analytical value if the staining conditions create uneven loading, excessive spread in the parent population, or unnecessary stress on the cells before the assay has even started. For proliferation analysis, the best staining condition is not necessarily the one that gives the brightest initial population. It is the one that produces a clean, reproducible starting point and preserves cell behavior well enough for later signal changes to remain interpretable.
- Key considerations before staining: Cell state before staining is often underestimated. Density, recovery status, recent handling, buffer composition, and baseline cell health can all affect how evenly the tracer is taken up and how reliably the population recovers afterward. A tracer loaded into a stressed or inconsistent sample may appear adequate at time zero but still produce poor-quality proliferation data later. This is why proliferation staining should be treated as a controlled analytical step rather than an incidental preparation stage.
- Staining intensity and signal balance: Strong fluorescence is not automatically better. If the initial staining is too aggressive, the signal may look impressive while compromising cell compatibility, broadening the starting distribution, or making later comparisons harder to interpret. If the signal is too weak, later generations may collapse into background. The practical objective is to create a parent population that is bright enough to remain distinguishable over the expected assay window, but narrow enough to preserve analytical separation as cells divide. This balance is much more important than maximizing absolute intensity.
- Fluorescent staining in multicolor assays: Proliferation tracer workflows are often embedded in more complex multicolor experiments, especially when researchers need to determine which phenotype-defined subpopulation is dividing. In those designs, tracer selection has to account for excitation source, emission overlap, relative brightness, and how strongly the tracer channel will compete with other readouts. A tracer that is acceptable in a single-color pilot may become a poor choice when the panel later includes phenotype markers, intracellular probes, or viability staining. Thoughtful fluorescent staining design therefore improves not only signal quality, but the interpretability of the assay as a whole.
- Integration with broader staining workflows: Proliferation tracing rarely exists in isolation. Many laboratories build it into broader workflows that already involve fluorescent reporters, organelle stains, viability channels, or structural labels. This is why users often consider proliferation tracers in parallel with more general fluorescent dyes and fluorescent probes, rather than treating them as a completely separate category. In practice, the proliferation tracer should integrate into the broader staining logic of the experiment without undermining either the tracer readout or the other channels it needs to coexist with.
Choosing the Right Cell Proliferation Tracer
The best proliferation tracer is the one that fits the scientific question, analytical platform, and staining constraints of the actual experiment. There is no universal best tracer because different assays place different demands on signal retention, spectral position, assay duration, and post-staining interpretation. Researchers therefore benefit more from a structured choice process than from choosing whichever reagent is most familiar. A rational selection strategy starts with what needs to be measured, then moves toward how the tracer needs to behave under those conditions.
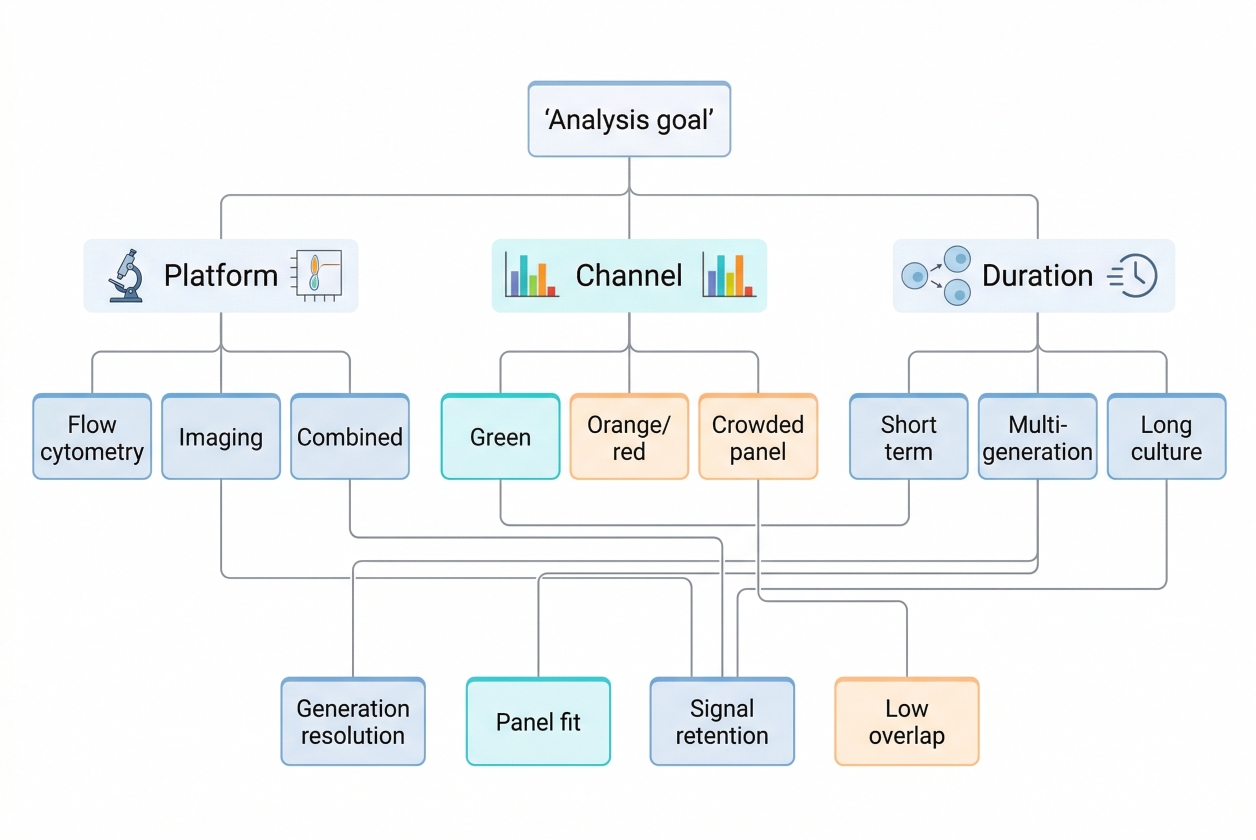 Fig. 2. Choose tracers by platform, panel, and assay duration (BOC Sciences Authorized).
Fig. 2. Choose tracers by platform, panel, and assay duration (BOC Sciences Authorized).
- Selection by detection platform: If the experiment is designed for population-level generation analysis, the tracer must support a signal distribution that can be read cleanly on the chosen analytical platform. Flow-cytometric workflows often prioritize clear parent-peak definition and generation separation, while imaging-linked workflows may place additional emphasis on spatial context and fluorescent stability in visual observation. The more quantitative the division-tracking requirement, the more important it becomes to choose a tracer whose signal behavior aligns with population-level analysis rather than simple fluorescent visibility.
- Selection by spectral requirements: Spectral planning matters early. Researchers should not ask only whether a tracer is green, orange, or blue, but whether that color occupies valuable panel space, whether the detector configuration supports it cleanly, and whether the tracer will interfere with phenotype markers or auxiliary readouts. A tracer assigned to a crowded channel may create more analysis burden than benefit. Spectral selection is therefore best treated as part of assay architecture, not as a cosmetic preference.
- Selection by cell type and assay duration: Cell type affects staining tolerance, division kinetics, and how quickly tracer resolution may compress over time. Fast-dividing populations may require stronger starting separation or shorter readout windows, while slowly dividing populations may place more value on stability and lower assay disturbance. The expected duration of the assay also matters because a tracer that works well over a short window may provide less useful discrimination after multiple rounds of culture. Selection should therefore account for both biological tempo and technical persistence.
- Selection by experimental goal: Some experiments need precise generational tracing. Others only need to distinguish dividing from non-dividing populations or to maintain a labeled cell population during a mixed-culture workflow. These goals are related but not identical. A tracer chosen for strict division-history analysis may differ from one selected for broader fluorescent cell tracking. The decision should therefore begin with the most important analytical question, not with the reagent list.
Building a proliferation-tracing workflow?
We can help compare tracer dyes, spectral options, and staining strategies to support cleaner proliferation data and better panel compatibility.
Experimental Design and Workflow Considerations
Good proliferation tracer data depend on more than tracer identity. Experimental design determines whether the fluorescent signal will be interpretable once cells begin to divide. For this reason, workflow planning should be treated as part of the assay logic rather than an afterthought. Controls, timing, acquisition consistency, and analysis discipline all influence whether a tracer experiment yields readable generation data or only a broad fluorescent shift with uncertain meaning.
- Sample preparation and control design: Well-prepared cells give the tracer a fair chance to perform as intended. Unstained controls help define baseline fluorescence, while single-stained controls help separate tracer signal from panel interactions in more complex experiments. In comparative studies, baseline population controls are especially important because they make it easier to determine whether apparent proliferation differences reflect real biology or differences introduced before the assay even started.
- Timing and division window: The timing of acquisition should match the expected division kinetics of the cell system. If cells have barely divided, the tracer may show little analytical structure beyond the parent population. If the assay runs too long, later generations may compress into low-intensity regions that are harder to separate cleanly. Choosing the correct window is therefore essential to preserving useful division information.
- Signal acquisition and analysis consistency: Once the assay moves into data collection, acquisition settings and analysis logic should remain consistent across groups. Changing detector settings, thresholds, or gating strategy between samples can create false differences that appear biological but are actually procedural. Proliferation tracer experiments are especially sensitive to this because interpretation depends heavily on distribution shape rather than signal presence alone.
- Workflow planning for reproducible results: Reproducibility improves when staining conditions, incubation timing, washing sequence, recovery interval, acquisition window, and analysis rules are documented together. A tracer assay is far more robust when these factors are planned as part of a defined cell staining protocol rather than improvised around the reagent alone.
Data Interpretation in Proliferation Tracer Experiments
Interpreting proliferation tracer data requires more discipline than simply looking for a weaker signal at a later time point. A reliable analysis asks whether the starting population was well defined, whether the signal distribution changed in a way consistent with division-linked dilution, and whether alternative explanations such as staining loss, panel conflict, or changing background can be ruled out. In other words, tracer interpretation is not only about what the fluorescence did, but about whether the way it changed remains consistent with the logic of the assay.
- Parent peak and generation structure: The parent population is the analytical anchor of the assay. If the initial signal distribution is too broad or poorly defined, later generations become difficult to resolve with confidence. A clean parent peak therefore matters not just at time zero, but across the entire interpretive framework of the experiment.
- Signal loss and true proliferation: Fluorescence reduction is only meaningful if it reflects cell division rather than dye leakage, poor retention, uneven loading, or changes in background. A dimmer population is not automatically a more proliferative population. The interpretation must remain tied to tracer behavior and experimental controls.
- Population heterogeneity in tracer data: One of the greatest advantages of proliferation tracers is that they can reveal heterogeneity within a cell population. Instead of collapsing the result into a single average signal, the tracer can show whether part of the population divided strongly while another fraction remained largely undivided. This adds much more interpretive depth than a total cell count alone.
- Combining tracer readouts with phenotypic markers: In many multicolor workflows, the most useful proliferation question is not simply whether cells divided, but which cells divided. Pairing tracer data with phenotype markers can reveal how proliferation behavior differs between subpopulations, treatment-defined states, or mixed-cell systems. This turns the tracer from a generic proliferation readout into a much more targeted analytical tool.
Common Problems and Troubleshooting
Problems in proliferation tracer experiments do not always appear as total assay failure. More often, the data look usable at first glance but carry hidden interpretive weakness because the staining was uneven, the readout window was poorly chosen, or the tracer channel became compromised inside a larger panel. Troubleshooting is therefore not only about increasing signal. It is about restoring the relationship between tracer behavior and biological meaning.
- Uneven initial staining: If the starting population is labeled unevenly, the parent peak becomes broad and later generations become harder to separate. This is one of the most common sources of downstream ambiguity because the assay may appear bright while still being structurally weak from the start.
- Poor peak resolution after several divisions: As divisions accumulate, signal compression can reduce the separation between generations. This effect becomes more severe when the initial staining distribution is already broad or when the readout is collected after too many rounds of division relative to the analytical sensitivity of the system.
- Toxicity and proliferation disturbance: A tracer is only useful if the cells remain able to behave in a representative way after staining. Excessive loading or non-ideal staining conditions can reduce proliferation, alter cell state, or introduce selection pressure that changes the very phenomenon being measured.
- Spectral overlap in multicolor panels: A tracer channel that looks acceptable alone may become much less reliable once phenotype markers, reporters, or other bright fluorophores are added. Compensation burden and channel interference can weaken the interpretability of the tracer even when the reagent itself is functioning correctly.
- Misinterpretation of weak signals: A weak signal should not automatically be interpreted as evidence of strong proliferation. In some experiments, the signal may be weak because of poor staining, loss during handling, or failure of the tracer to remain associated with the cells under the chosen conditions. Controls and analysis logic are essential for distinguishing these possibilities.
Research Uses of Cell Proliferation Tracers
Cell proliferation tracers are valuable because they do more than indicate whether a population has expanded. They help researchers understand how proliferation is distributed across a sample, whether only a subset of cells responds to stimulation, how division behavior evolves over time, and how proliferative activity relates to phenotype, treatment condition, or experimental context. In many workflows, this level of resolution is essential because total cell number alone can hide important biological differences. Two samples may show similar final cell counts while still having very different division histories, different fractions of responding cells, or different relationships between proliferation and cell identity. That is why proliferation tracers remain highly useful in experiments that require population-level structure, temporal interpretation, and integration with broader fluorescent analysis strategies.
Immune Cell Proliferation Studies
One of the most established uses of cell proliferation tracers is in immune-cell proliferation analysis, where researchers often need to determine how strongly a defined lymphocyte or mixed immune population responds to stimulation. In these workflows, total expansion alone is rarely enough. What often matters more is whether the entire population enters division, whether only a responsive subfraction expands, how many generations are reached within the assay window, and whether that proliferative behavior differs between donor groups, treatment conditions, or stimulation protocols. A proliferation tracer is especially useful here because it allows immune-cell responses to be interpreted as structured generational distributions rather than as bulk signal changes. This becomes even more informative when the tracer is paired with phenotype markers, because the experiment can then distinguish not only whether proliferation occurred, but which immune subsets were responsible for it and whether proliferative intensity was evenly distributed across those subsets.
Mixed-Population and Co-Culture Assays
Cell proliferation tracers are also highly valuable in mixed-population and co-culture workflows, where the main analytical challenge is often not measuring proliferation in isolation but assigning proliferation behavior to the correct cell population within a more complex system. In these assays, multiple cell types may influence one another through contact, soluble factors, matrix effects, or competition for nutrients, making total cell number a poor substitute for true population-level interpretation. A tracer allows one pre-labeled population to be followed over time so that researchers can determine whether it continues to divide, whether its proliferation is suppressed or enhanced in co-culture, and whether those changes occur uniformly or only within a subset of labeled cells. This is particularly important when one population remains unlabeled or is identified through separate markers, because the tracer adds temporal structure to what would otherwise be a static mixed-cell readout. In well-designed co-culture experiments, proliferation tracers therefore help separate cell identity, cell interaction, and division behavior into interpretable layers rather than collapsing them into a single endpoint measurement.
Phenotype-Linked Proliferation Analysis
In many research settings, the most informative proliferation question is not simply whether cells are dividing, but which cells are dividing and how proliferative behavior aligns with phenotype. This is where phenotype-linked proliferation analysis becomes especially powerful. By combining a proliferation tracer with surface markers, intracellular markers, reporter signals, or state-defining fluorescence channels, researchers can move from a generic proliferation assay to a much more specific biological interpretation. For example, the experiment can distinguish whether proliferating cells belong to one phenotypic fraction or another, whether proliferative activity correlates with activation state, whether proliferative capacity shifts after treatment, or whether a phenotype-defined subgroup remains largely undivided even when the broader population expands. This type of analysis is particularly valuable in heterogeneous populations because it prevents the average proliferation signal from masking important subgroup behavior. From a workflow perspective, phenotype-linked proliferation studies demand more careful panel design and compensation planning, but the increase in information density is often substantial enough to justify that added complexity.
Longitudinal Tracking in Cell-Based Studies
Longitudinal studies benefit from proliferation tracers because these reagents make it possible to follow how a labeled population changes over a defined experimental window rather than relying on a single terminal readout. In this context, the tracer is useful not only for identifying whether proliferation occurred, but for showing how the pace and structure of division develop across time. Researchers can compare early versus late proliferative behavior, determine whether the population continues to divide steadily or plateaus after an initial burst, and assess whether later cultures become increasingly heterogeneous as some cells continue cycling while others stop responding. This kind of time-aware interpretation is especially important in experiments involving repeated stimulation, extended culture, recovery phases, or progressive environmental change, because proliferation may not follow a simple monotonic pattern. The tracer therefore adds a historical dimension to the assay, helping researchers see whether the final state of the culture reflects continuous expansion, selective outgrowth, transient response, or gradual divergence into multiple proliferative states.
How BOC Sciences Supports Cell Proliferation Tracer Workflows?
BOC Sciences supports cell proliferation tracer workflows through a combination of standard product supply, custom synthesis, application-focused technical support, and quality documentation services. Our goal is not only to provide fluorescent reagents, but to help researchers align tracer selection with assay design, analytical platform, cell type, and workflow scale. Because cell proliferation tracing often depends on the balance between signal retention, spectral compatibility, cell loading behavior, and downstream readout quality, practical support should extend beyond reagent availability alone. BOC Sciences therefore provides both ready-to-use options and more tailored solutions for laboratories developing, optimizing, or scaling proliferation-tracking assays.
Ready-To-Use Proliferation Tracer Products
- Supply of multiple classic cell proliferation tracer dyes, including CFSE-type reagents and related fluorescent tracing compounds for division-linked staining workflows.
- Coverage of multiple fluorescence wavelength options to support different detection platforms, multicolor panel layouts, and instrument configurations.
- High-purity and high-stability product options suitable for routine staining, assay development, and comparative proliferation analysis.
- Multiple package sizes available to match different research scales, from early-stage method development to larger experimental programs.
Custom Proliferation Tracer Design And Synthesis
- Custom design of cell proliferation tracer structures according to intended staining route, signal logic, and experimental requirements.
- Support for multiple fluorophore choices, including FITC, Cy-series, Rhodamine, and other fluorescence scaffolds for tailored tracer development.
- Functional modification options to improve properties such as cell permeability, labeling performance, or workflow-specific targeting behavior.
- Synthesis support covering early research, method optimization, and scale-up stages for projects that require more than off-the-shelf tracer selection.
Workflow Optimization And Technical Guidance
- Practical support for optimizing cell labeling and proliferation detection strategies according to assay goal, expected division window, and analytical format.
- Guidance for both flow cytometry-based proliferation analysis and fluorescence imaging-related workflows where tracer performance must remain interpretable.
- Staining recommendations adapted to different cell types, helping users improve loading consistency and reduce workflow mismatch.
- Troubleshooting support for common challenges such as signal attenuation, weak generation resolution, staining imbalance, and tracer-associated cytotoxicity concerns.
Quality Documentation And Compliance Support
- Complete quality testing support with COA documentation to improve traceability and experimental confidence.
- Support for customized purity specifications and project-specific quality requirements where tighter control standards are needed.
- Stability-related information and storage recommendations to help preserve reagent performance during transport, handling, and routine use.
- Availability of compliance-related documents such as SDS files for laboratories requiring structured documentation support.
Do You Need A Consultation?
BOC Sciences integrates cutting-edge fluorescence technologies to accelerate your research, driving next-generation solutions for drug discovery and diagnostics.
Transform Your Studies with Cutting-Edge Fluorescent Products
| Catalog | Name | CAS | Inquiry |
|---|---|---|---|
| A16-0032 | CytoTrace™ Orange CMTMR | 323192-14-9 | Bulk Inquiry |
| A16-0175 | Lucifer Yellow CH Dipotassium Salt | 71206-95-6 | Bulk Inquiry |
| A16-0070 | Biocytin hydrazide | 102743-85-1 | Bulk Inquiry |
| A16-0025 | 5(6)-Carboxyfluorescein diacetate | 124387-19-5 | Bulk Inquiry |
| A16-0081 | 6-Carboxyfluorescein diacetate succinimidyl ester | 150206-15-8 | Bulk Inquiry |
| A16-0024 | Green CMFDA | 136832-63-8 | Bulk Inquiry |
| A16-0091 | Lucifer yellow CH ammonium salt | 188904-20-3 | Bulk Inquiry |
| A16-0067 | 6-Carboxyfluorescein diacetate | 3348-03-6 | Bulk Inquiry |
| A16-0078 | Lucifer yellow cadaverine | 149733-79-9 | Bulk Inquiry |
| A16-0077 | 7-Amino-4-chloromethylcoumarin | 147963-22-2 | Bulk Inquiry |
High-Performance Fluorescent Tools for Your Research
- Nitric Oxide (NO) & Reactive Oxygen Species (ROS) Probes for oxidative stress and signaling detection.
- Nuclear Fluorescent Probes DNA-binding dyes for nucleus visualization.
- Lipid Fluorescent Probes Dyes for lipid droplets and membranes.
- Fluorescent Probes Versatile tools for biomolecular and cellular imaging.
- Lysosomal Fluorescent Probes Acidic organelle markers for lysosome tracking.
- Calcium, Chloride and Other indicators Fluorescent indicators for intracellular ion flux monitoring.
- pH Indicators Fluorescent sensors for intracellular pH monitoring.
- Metal Fluorescent Probes Selective sensors for intracellular metal ions.
Explore More Topics
Online Inquiry