Cell Staining: Principles, Methods, Dyes, and Practical Selection Guide
Cell staining is one of the most widely used approaches in cell analysis because it helps researchers make otherwise hard-to-distinguish cellular structures easier to observe, compare, and interpret. In fluorescence-based workflows, staining is not only about making cells visible under a microscope. It is also about deciding what kind of information the experiment is expected to generate. In some cases, the goal is to reveal overall morphology. In others, the priority is to highlight a specific organelle, distinguish living from damaged cells, compare treatment groups, or build a multicolor imaging panel that combines several readouts in one experiment. This is why a practical cell staining guide needs to do more than define the term. It should help readers connect staining principles with dye choice, protocol design, workflow decisions, and result interpretation.
This page is designed as a broad resource hub for cell staining. It introduces the core concepts behind staining, explains the main principles that influence signal quality, and outlines how fluorescent dyes, protocol design, and application goals fit together. At the same time, some important questions deserve deeper treatment than a single overview page can provide. If you are deciding between live cell staining and fixed cell staining, building a cell staining protocol, selecting a fluorescent dye for cell staining, planning multicolor cell staining, or exploring organelle-specific cell staining, the related topic pages linked throughout this article provide more focused guidance.
Cell Staining Overview: What This Guide Covers
Cell staining is a broad topic, and the right reading path often depends on the experiment you are trying to design. Some users need a foundational overview of what staining is and why it matters. Others already understand the basics and need practical guidance on selecting dyes, minimizing background, choosing between live and fixed workflows, or deciding which organelle should be labeled first. For that reason, this page works as a central pillar rather than a narrow technical note. It introduces the main staining concepts while also helping readers navigate to more specific topics when the experimental question becomes more defined.
- Start here for the basics: If you want a high-level understanding of what cell staining is, why it is used, and what kinds of information it can reveal, the next sections provide a practical overview of definition, principles, common methods, and fluorescent dye logic.
- Go deeper into workflow decisions: If your main challenge is choosing a staining strategy, compare live cell staining vs fixed cell staining, review a more detailed cell staining protocol, or explore common problems in cell staining and how to fix them.
- Choose dyes more rationally: If the staining question is mainly about reagent selection, spectral fit, or assay compatibility, the most useful next step is often how to choose the right fluorescent dye for cell staining or a broader guide to multicolor cell staining.
- Navigate by biological target: If your experiment is centered on one structure or organelle, such as the nucleus, mitochondria, lysosomes, membrane, endoplasmic reticulum, Golgi apparatus, or cytoskeleton, the organelle-focused pages linked later in this guide will usually provide a better decision framework than a general overview alone.
What Is Cell Staining and Why Is It Important?
The cell staining is a critical experimental method used in biology and medicine. It wants to enchant cells and their organelles with dyes or markings so they can be easier to see under a microscope. This process starts with the dye and the cells that make it. The dye attaches to many intracellular structures and creates seen complexes describing the cell's structures and processes. Researchers fix samples when they do cell staining, so that the cells' structures are preserved. Then the right dyes are chosen that bind to the right molecules in the cells (like nucleic acids, proteins or lipids). Not only can stains enable us to see the change of the cells' appearance, but they can also reveal their activity through their morphological changes, including cell division, death and signal transduction. Each dye, thanks to its unique binding ability to cellular proteins, can be assigned to individual organs of the cell, so that scientists can see complex structure within the cell with a microscope. The use of cell staining is endless, from basic biology to clinical pathology, whose very existence depends on it.
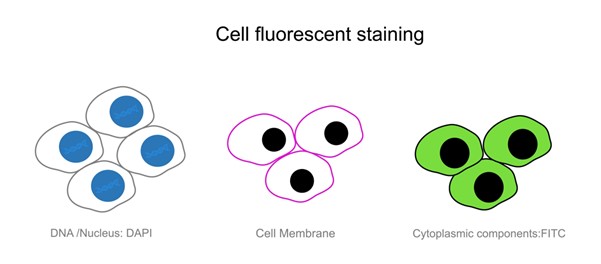 Fig. 1. Live cell staining (BOC Sciences Authorized).
Fig. 1. Live cell staining (BOC Sciences Authorized).
The importance of cell staining comes from the limits of direct observation. Many cells, especially under standard brightfield conditions, provide only limited visual contrast. Even when cell outlines are visible, internal structures may remain poorly resolved. Staining helps translate biological organization into a signal that can be recognized under the imaging method being used. In fluorescence imaging, this usually means converting a target structure into a defined optical channel with enough contrast to support comparison. That comparison may be as simple as determining where the nucleus is located, or as complex as assessing whether different organelles can be distinguished cleanly in a multicolor panel.
Core Principles of Cell Staining
It's all about the chemical bonds between the dye and the cells that drive the science of cell staining. The dye attaches to intracellular molecules (including nucleic acids, proteins and lipids) in visualized complexes that both recognise and reveal internal cells. The chemical features and compatibility with the molecules that they are applied to determine which dyes are chosen and used. Dye binding can happen in a number of different ways. There is electrostatic attraction, for which many cationic dyes are drawn to anionic proteins in the cell membrane or to DNA and RNA in the interior of the cell, so that the dye could get inside and stick to the target molecule. Also of note is hydrogen bonding and hydrophobic interaction. These reactions induce dye molecules to attach to intracellular structures, which increases staining specificity and efficiency. What's more, staining conditions like pH, temperature, stain time and so on can alter the outcomes. It can be easier to bind dyes to cell elements if conditions are right and target molecules in the cell are more easily stained. Scientists usually fine-tune these conditions according to experimental demands in order to get the best staining. In addition to being direct morphological, cell staining reveals biochemical activity within cells, all of which are important insights for future biological study through these principles and processes.
- Target interaction determines meaning: Some dyes bind preferentially to nucleic acids, some partition into membranes, some accumulate in specific organelles, and some are chosen because their signal is compatible with a larger assay rather than because they are universally optimal. The important point is that signal meaning comes from the interaction logic of the stain, not from brightness alone.
- Specificity and background are inseparable: A strong signal is only useful when it can be distinguished from non-specific fluorescence, diffuse labeling, or channel interference. Many staining problems arise because a visually bright image is mistaken for a selective one. In practice, the most useful stain is often the one that produces the clearest interpretable contrast rather than the highest apparent intensity.
- Sample handling changes staining behavior: Cell density, medium composition, incubation time, temperature, wash stringency, and delay before imaging can all affect the way fluorescence appears. These variables do not just influence signal strength. They can alter the balance between target-associated staining and background, which in turn changes how the image should be interpreted.
- Optical context matters: The same dye may be easy to use in a single-channel workflow and much harder to manage in a multicolor design. Excitation and emission placement, instrument configuration, exposure settings, and crosstalk risk should all be considered part of staining logic, not secondary considerations added after the stain has already been chosen.
How to Choose the Right Cell Staining Strategy?
Choosing a staining strategy starts with the experimental question. Before comparing products or selecting colors, it is helpful to decide what kind of readout the experiment truly needs. A structural localization experiment, a viability-oriented assay, a tracking workflow, and a multicolor organelle panel are all cell staining applications, but they do not demand the same staining logic. The most reliable selection process therefore begins with purpose, then moves to sample state, then to dye properties and imaging constraints.
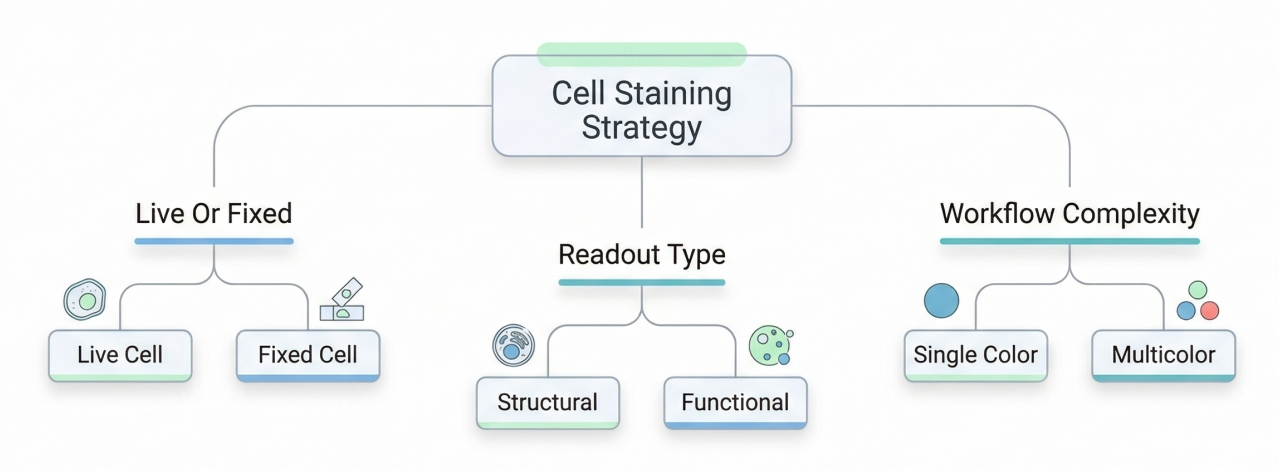 Fig. 2. Choosing a staining strategy depends on sample and readout (BOC Sciences Authorized).
Fig. 2. Choosing a staining strategy depends on sample and readout (BOC Sciences Authorized).
Live Cell Staining vs Fixed Cell Staining
One of the most common strategic choices is whether the stain should be applied in living cells or in a fixed sample. Live cell staining vs fixed cell staining is not simply a matter of convenience. Live-cell workflows are more suitable when temporal behavior, immediate localization, or short-term dynamic observation matters. Fixed-cell workflows are often preferred when the experiment requires greater processing flexibility, longer storage stability, or post-treatment handling that would be difficult to maintain in living cells. The right choice depends on whether preserving a live-state readout is more important than gaining a more controlled endpoint workflow.
Structural Staining vs Viability Staining
Not all cell stains are intended to answer the same kind of question. Some are mainly used to reveal where structures are located, while others are chosen because the staining result supports a function-related or condition-related readout. This distinction is important because it affects how signal changes should be interpreted. A structural dye may help define organelle position or membrane boundaries, while a viability stain is intended to help distinguish live and dead cells under an assay framework designed for that purpose. Users working on this type of question should usually review cell viability staining separately from general structural imaging.
Single-Color Workflows vs Multicolor Workflows
A simple single-color experiment may allow the user to prioritize brightness and ease of detection. By contrast, a multicolor panel requires a more balanced decision process. The staining channel must fit together with the rest of the assay, not dominate it. This means that spectral spacing, channel hierarchy, expected signal strength, and image interpretation all need to be planned together. For users combining nucleus, membrane, and organelle markers in the same experiment, multicolor cell staining deserves its own design process rather than being treated as a routine extension of a single-dye assay.
Choosing between dyes, targets, and staining workflows?
We can help match cell staining reagents to your target structure, imaging setup, and workflow goals, from routine labeling to multicolor assay design.
Common Cell Staining Methods and Basic Protocol Concepts
Cell staining includes a wide range of traditional and fluorescence-based methods, and different methods were developed to answer different observational questions. Some older approaches remain important because they illustrate the logic of differential staining and contrast generation. Others are used less as direct models for fluorescence imaging but still provide conceptual value in understanding why one component stains differently from another. For that reason, a broad overview of common staining methods remains useful even on a fluorescence-oriented resource page.
Gram Staining
The Danish biologist Christian Gram in 1884 based his Gram staining method on the structure and chemical make-up of bacterial cell walls. It's a method that separates two classes of bacteria, Gram-positive and Gram-negative, based on how much they stain or fade away. Gram-positive bacteria are purple; Gram-negative bacteria are red after staining. This approach is transparent for the morphology, configuration and some structural features of bacterial organisms and is a key method for bacterial identification and classification.
Wright Staining
Wright's stain is mostly made of thiazine colours and eosin, which give the nucleus and cytoplasm various colours. When the nucleus is stained, it will often be blue-purple or reddish-purple and the cytoplasm can be blue or red. That hue difference allows you to make distinct parts of the cell with a microscope. The Wright staining is mostly applied in hematological analyses like blood smears and bone marrow smears. It is very convenient to see all kinds of white blood cells, red blood cells and nucleated red blood cells. It is also employed for the analysis of certain microbes such as trichomonas and fungi. Wright staining, which offers sharp images of the cellular interior, especially of the nucleus and cytoplasm, is one of the most common types of staining in clinical labs, and for all sorts of samples.
Giemsa Staining
Giemsa staining relies on interactions between proteins and dyes. Basic fuchsin binds to positively charged DNA, while acidic eosin binds to negatively charged proteins. These specific charge interactions allow the dye molecules to embed into the cell structures, achieving a staining effect. Giemsa staining offers excellent contrast between the nucleus and cytoplasm, helping to clearly differentiate various cell types. It is primarily used for staining blood smears, tissue sections, and microbial specimens. This method is particularly useful for observing Plasmodium, sperm acrosomes, and other small structures.
Wright-Giemsa Staining
Wright-Giemsa staining is a standard technique of stains on cells and tissues, usually for clinical diagnosis and teaching. It is a combination of Wright and Giemsa stain with two different dyes to add extra stains. This rule depends on the affinity differences between the dyes and cellular parts. Acid dyes like eosin stick mostly to proteins in the cytoplasm, and basic dyes like azure stick to DNA in the nucleus. Such binding selectively means nucleus is distinct from cytoplasm. Wright-Giemsa staining is used in many biological studies and clinical diagnostics, particularly in hematology, pathology and microbiology. It shows detailed cell-structure photos which helps in cell-type identification and classification.
Papanicolaou Staining
Papanicolaou staining (also known as Pap staining) is a very popular cytological staining method used to color exfoliated cell smears. The best thing about this staining is the bright, sharp colors of the staining that make it easy to see cell features. For pap staining, the dye used usually uses the EA36 (has hematoxylin and orange G as components). In modified versions, original parts might be removed, and different dyes substituted to optimise the staining. Pap staining is based on the different preference of cells and tissues for dyes. It's this differential affinity that enables cells (the nucleus and the cytoplasm) to be marked with dyes of specific colours under the microscope to make it possible to distinguish cell types and disease states. Pap is now standard in Gynaecological cytology, thyroid fine-needle aspiration cytology and other exfoliated cell diagnostics. Particularly, it is ideal for cervical cancer diagnosis and the detection of other cancers early.
HE Staining
The staining technique Hematoxylin and Eosin (H&E) is an old staining method widely employed in pathology, histology, and biology. There are two major dyes employed here, hematoxylin (for staining the cell nucleus) and eosin (for staining the cytoplasm). The principle of H&E consists of different affinities of hematoxylin and eosin towards hydrophilic and lipophilic molecules so that the different cell types can be distinguished. Hematoxylin reaches down to the nucleus and tethers to DNA, while eosin is mainly tethered to proteins and non-lipid molecules in the cytoplasm. H&E staining is most often employed to observe cell shape and size in sections of tissue and it's a core diagnostic tool in clinical pathology. It can identify tissues changes in various disease conditions like inflammation and tumors.
General Workflow for Reliable Cell Staining Results
Reliable cell staining depends on disciplined workflow design. Many staining issues arise because the reagent is selected first and the experimental logic is defined later. A better approach is to align sample state, staining conditions, imaging schedule, and controls before drawing conclusions from the fluorescence readout. The goal is not just to obtain a visible image, but to obtain one that remains interpretable across replicate samples and comparison groups.
- Prepare cells according to the intended readout: Before staining begins, define whether the experiment is meant to emphasize morphology, organelle localization, viability, proliferation, or cell tracking. Cell density, attachment state, treatment timing, and handling sequence should all be compatible with that goal.
- Match dye loading to actual workflow conditions: Concentration, incubation time, and wash steps should be evaluated together rather than one at a time. A condition that looks acceptable in a pilot image may still perform poorly across the full acquisition workflow if imaging is delayed or if background rises after washing.
- Control timing and acquisition settings: Staining quality is influenced by when the sample is imaged and under what settings. Exposure, gain, filter configuration, and channel order can all influence how the stain appears. These variables should be documented as part of the staining workflow, not only as part of the imaging workflow.
- Use controls to improve confidence: Unstained samples, single-stain controls, and matched treatment groups are often essential for distinguishing true staining behavior from background or acquisition drift. This is especially important when results will be compared between groups or integrated into a multicolor design.
Users who want a fuller execution framework, including workflow sequencing and troubleshooting logic, should continue to the dedicated guide on reliable cell staining protocols. When the main challenge is weak signal, high background, diffuse labeling, or inconsistency between repeats, the next most useful page is often common problems in cell staining and how to fix them.
List of Conventional Morphological Cell Stains
In the staining process, the choice of dyes is crucial for achieving the desired staining effect. Different dyes have varying specificities. For example, some dyes, like DAPI and Hoechst, have a high affinity for DNA, allowing them to specifically bind to DNA and cause the nucleus to fluoresce, making it easier to observe under a fluorescence microscope. Other dyes, such as fluorescein and crystal violet, are often used to label the cell membrane or cytoplasm, as they can effectively bind to specific components on the cell membrane, helping researchers observe cell morphology and structure.
| Dyes | Applications |
| Congo Red | Congo red is an acidic, dark red powder dye, soluble in water and alcohol, turning blue in acidic conditions. It is used as a counterstain for hematoxylin in plant sections and can stain colloids or cellulose red in the cytoplasm. It is also used for staining animal tissues, including nerve axons, elastic fibers, and embryonic materials. Due to its solubility in both water and alcohol, quick washing and dehydration are necessary. |
| Methyl Blue | Methyl blue is a weak acidic dye, soluble in water and alcohol. It is widely used in both animal and plant section techniques. Methyl blue, when combined with eosin, is used to stain nerve cells and is an essential dye in bacterial preparations. Its aqueous solution is a live stain for protozoa, though it oxidizes easily and thus stained samples cannot be preserved long term. |
| Eosin | Eosin Y is an acidic dye in the form of reddish-blue small crystals or brown powder, soluble in water (with a solubility of 44% at 15°C) and alcohol (solubility in anhydrous alcohol is 2%). Eosin is widely used in animal tissue preparations as an excellent cytoplasm dye and is often used as a counterstain for hematoxylin. |
| Acid Fuchsin | This is an acidic dye in red powder form, soluble in water and slightly soluble in alcohol (0.3%). It is widely used in animal tissue preparations and in plants to stain parenchymal cells and cellulose walls. When combined with methyl green, it can show mitochondria. |
| Fast Green | Fast green is an acidic dye, soluble in water (solubility of 4%) and alcohol (solubility of 9%). It is used extensively for staining cellulose-containing tissues in plants and cells. |
| Aniline Blue | Aniline blue is a mixed acidic dye, typically lacking a fixed standard. It is used in plant section techniques, often in combination with safranin for tissue staining, and can also be used for staining algae. |
| Sudan III | Sudan III is a weak acidic dye that does not dissolve in water, appearing as a red powder and readily dissolving in fats and alcohol (solubility of 0.15%). It is used as a fat stain. |
| Sudan IV | Sudan IV is also a weak acidic dye and an excellent fat stain, now commonly used to replace Sudan III. It can stain resins, milk ducts, wax, and keratin structures, and can also stain chloroplasts dark red. |
| Basic Fuchsin | Basic fuchsin is a basic dye in dark red powder or crystal form, soluble in water (solubility of 1%) and alcohol (solubility of 8%). It is used to stain collagen fibers, elastic fibers, granules in eosinophils, and the nuclear substance of central nervous tissues. It is also used to identify Mycobacterium tuberculosis. |
| Crystal Violet | Crystal violet is a basic dye, soluble in water (solubility of 9%) and alcohol (solubility of 8.75%). It is widely used in cytology, histology, and bacteriology as an excellent nuclear stain, often used to visualize centrosomes of chromosomes, and can stain starch, fibrin, neuroglia, and other structures. |
| Gentian Violet | Gentian violet is a mixed basic dye, primarily consisting of crystal violet and methyl violet. When necessary, it can substitute for crystal violet. In medicine, it is used as a component of violet antiseptic solution, and methyl violet can substitute for both gentian violet and crystal violet when needed. |
| Neutral Red | Neutral red is a weak basic dye in red powder form, soluble in water (solubility of 4%) and alcohol (solubility of 1.8%). It is commonly used as a live stain for protozoa and to display the internal contents of live cells in plant and animal tissues. Aged aqueous solutions of neutral red are frequently used to display Nissl bodies. |
Fluorescent Dyes for Cell Staining and How to Choose Them?
Fluorescent staining techniques are essential tools for studying cellular structure and function. By labeling specific cellular structures and using fluorescence microscopy, researchers can reveal intricate processes within cells. Below are common fluorescent staining techniques for various cellular components, including the cell membrane, cytoskeleton, endoplasmic reticulum, Golgi apparatus, lipids, lysosomes, mitochondria, and nucleus, along with their applications.
The cell membrane acts as a barrier between the cell and its environment, regulating material exchange and signal transmission. Common dyes such as DiI and DIO are lipid-soluble fluorescent stains that embed into the bilayer of the cell membrane, providing long-term membrane labeling. DiI emits red fluorescence, while DIO emits green fluorescence, both widely used in studies on cell migration, membrane dynamics, and cell fusion. Phospholipids tagged with fluorophores are also used in membrane imaging, particularly for analyzing membrane fluidity and structural integrity.
| Cat. No. | Product Name | CAS No. | Inquiry |
| A16-0086 | 1,6-Diphenyl-1,3,5-hexatriene | 1720-32-7 | Inquiry |
| A16-0171 | Octadecyl Rhodamine B Chloride | 65603-19-2 | Inquiry |
| A16-0008 | DiD perchlorate | 127274-91-3 | Inquiry |
| A16-0035 | Calcein Blue AM | 168482-84-6 | Inquiry |
| F02-0108 | Cy3 dic18 | 22366-93-4 | Inquiry |
| F02-0115 | Cy5 dic18 | 75539-51-4 | Inquiry |
The cytoskeleton consists of microtubules, microfilaments, and intermediate filaments, maintaining cell shape, providing mechanical support, and participating in cell movement and division. Phalloidin is a classic tool for staining microfilaments (F-actin), as it binds to F-actin with high affinity, revealing their distribution within cells. Phalloidin is often conjugated to fluorophores like Alexa Fluor or FITC for imaging. Anti-α-tubulin antibodies are commonly used for microtubule staining, with secondary antibodies (e.g., Alexa Fluor 488) providing clear visualization of the microtubule network. For intermediate filaments, specific antibodies such as cytokeratin or vimentin antibodies are used to label corresponding filament structures.
| Cat. No. | Product Name | CAS No. | Inquiry |
| A16-0004 | Phalloidin-FITC | 915026-99-2 | Inquiry |
| A16-0106 | BF 568 Phalloidin | N/A | Inquiry |
| A16-0111 | BF 594 Phalloidin | 330626-83-0 | Inquiry |
| A16-0100 | BF 350 Phalloidin | N/A | Inquiry |
| A16-0002 | Phalloidin-TRITC | 915013-10-4 | Inquiry |
| A16-0003 | Phalloidin-TFAX 488 | 289620-19-5 | Inquiry |
The endoplasmic reticulum (ER) is the site of protein synthesis, lipid metabolism, and calcium storage. ER-Tracker dyes are highly efficient for ER-specific staining, interacting with components of the ER membrane to emit fluorescence. Common ER-Tracker dyes include ER-Tracker Green and ER-Tracker Red, which emit green and red fluorescence, respectively, and are suitable for both live and fixed cells. ER staining is primarily used in studies of protein folding, ER stress, and intracellular calcium signaling pathways.
| Cat. No. | Product Name | CAS No. | Inquiry |
| A16-0096 | ER-Tracker Blue-White DPX | N/A | Inquiry |
| A16-0062 | IraZolve-L1™ | 2169684-98-2 | Inquiry |
| A16-0064 | ReZolve-ER™ | 1404104-40-0 | Inquiry |
| A16-0048 | 3,3'-Dihexyloxacarbocyanine iodide | 53213-82-4 | Inquiry |
| A16-0214 | LumiTracker ER Green | N/A | Inquiry |
| A16-0215 | LumiTracker ER Red | N/A | Inquiry |
The Golgi apparatus is responsible for modifying, sorting, and transporting proteins and lipids. BODIPY FL C5-ceramide is a widely used Golgi stain that binds to Golgi lipids and emits green fluorescence. NBD C6-ceramide is another common lipid marker for the Golgi. Golgi staining reveals the complex morphology of this organelle, aiding in research on protein secretion pathways and intracellular transport. Additionally, antibodies targeting Golgi-specific proteins, such as GM130, can be used with fluorescent secondary antibodies for staining.
| Cat. No. | Product Name | CAS No. | Inquiry |
| A16-0059 | C6 NBD Ceramide | 94885-02-6 | Inquiry |
| A16-0186 | NBD C6-Ceramide | 86701-10-2 | Inquiry |
Lipids play key roles in cell signaling, membrane structure, and energy storage. Nile Red is a lipid-specific fluorescent dye that marks neutral lipids such as lipid droplets, emitting yellow or red fluorescence, making it a common tool in studies on lipid metabolism and adipocyte research. BODIPY dyes are another class of fluorescent lipid stains, known for selectively labeling fatty acids, cholesterol, and other lipid molecules, emitting strong green or red fluorescence. Lipid staining is crucial for investigating lipid metabolism disorders and associated disease mechanisms.
| Cat. No. | Product Name | CAS No. | Inquiry |
| A16-0020 | Laurdan | 74515-25-6 | Inquiry |
| A16-0044 | C6 NBD Sphingomyelin | 94885-04-8 | Inquiry |
| A16-0047 | C6 NBD Glucosylceramide | 94885-03-7 | Inquiry |
| A16-0144 | Nile Red | 7385-67-3 | Inquiry |
| A16-0073 | NBD Dihexadecylamine | 117056-66-3 | Inquiry |
| A16-0153 | NBD cholesterol | 78949-95-8 | Inquiry |
Mitochondria are the powerhouse of the cell, central to energy metabolism. MitoTracker dyes specifically label mitochondria in live cells with durable staining effects. MitoTracker Red and MitoTracker Green are commonly used dyes that emit red and green fluorescence, respectively. JC-1 dye is another mitochondrial stain used to assess mitochondrial membrane potential; it emits red fluorescence under high membrane potential and green when the membrane potential decreases. Mitochondrial fluorescence staining plays a key role in studies on energy metabolism, apoptosis, and mitochondrial diseases.
| Cat. No. | Product Name | CAS No. | Inquiry |
| A16-0159 | HIDC iodide | 36536-22-8 | Inquiry |
| A16-0170 | Rhodamine-123 | 62669-70-9 | Inquiry |
| A16-0093 | Rhodamine 6G | 989-38-8 | Inquiry |
| A16-0026 | Di-8-ANEPPS | 157134-53-7 | Inquiry |
| A16-0007 | MitoMark Red I | 167095-09-2 | Inquiry |
| A16-0104 | JC-9 Dye | 522592-13-8 | Inquiry |
The nucleus is the cell's storage center for genetic information, hosting functions such as chromosome arrangement and gene expression. DAPI is the most commonly used nuclear dye, binding to A-T-rich regions of DNA and emitting blue fluorescence. It is suitable for staining DNA in fixed cells or tissue sections. Hoechst dyes are another class of nuclear stains, emitting blue fluorescence and applicable to both live and fixed cells. Additionally, fluorescent-labeled antibodies can be used to stain specific nuclear proteins, such as nucleolar or nuclear membrane proteins.
| Cat. No. | Product Name | CAS No. | Inquiry |
| A16-0201 | DAPI dihydrochloride | 28718-90-3 | Inquiry |
| A16-0023 | 7-ethoxy-4-Methylcoumarin | 87-05-8 | Inquiry |
| A16-0223 | YO-PRO 3 | 157199-62-7 | Inquiry |
| A16-0222 | YOYO 1 | 143413-85-8 | Inquiry |
| A16-0220 | TOTO 3 | 166196-17-4 | Inquiry |
| A16-0221 | TO-PRO 1 | 157199-59-2 | Inquiry |
Cell Staining by Target Organelle
One of the most practical ways to organize cell staining decisions is by target organelle. Different intracellular structures demand different staining logic because they differ in accessibility, morphology, localization pattern, and imaging purpose. A nucleus stain is usually expected to behave differently from a membrane stain, and both differ from the design considerations associated with mitochondrial or lysosomal labeling. For users working with one main target, organelle-focused pages often provide more directly actionable guidance than a broad overview alone.
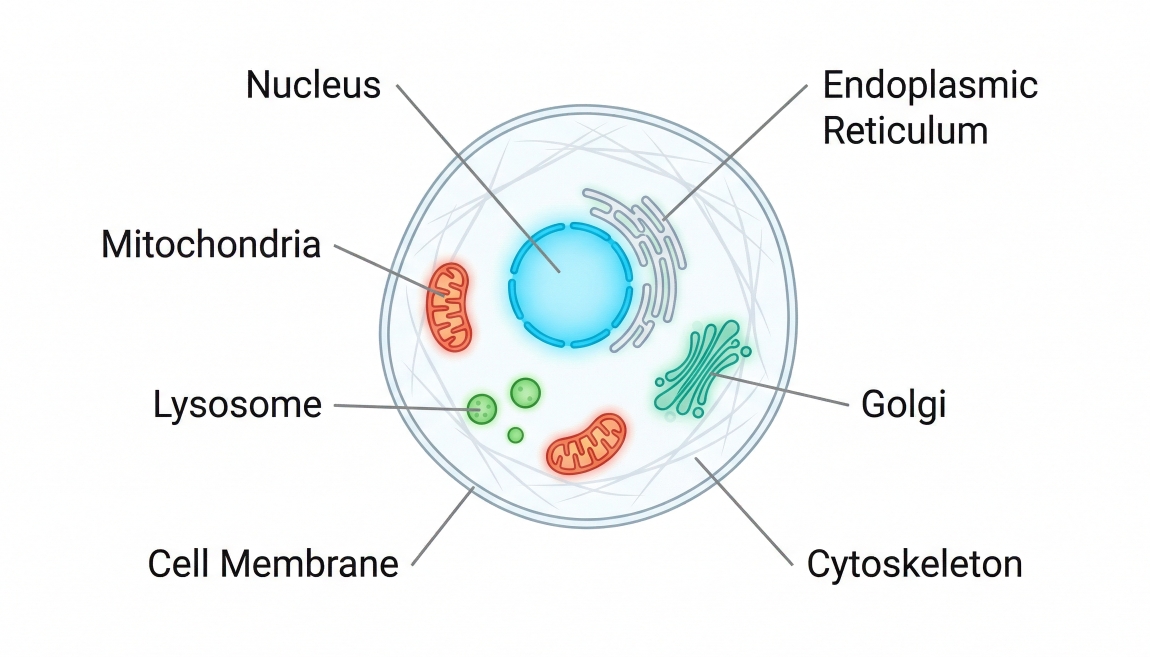 Fig. 3. Different targets require different cell staining approaches (BOC Sciences Authorized).
Fig. 3. Different targets require different cell staining approaches (BOC Sciences Authorized).
- Nucleus: Nuclear labeling is commonly used for cell counting, morphology support, segmentation, and spatial reference within multicolor assays. For a deeper framework, see a practical guide to nuclear staining in cells.
- Mitochondria: Mitochondrial staining can be used to examine organelle localization, network-like pattern, and in some workflows condition-linked behavior. For more detail, see mitochondrial staining in cells.
- Lysosomes: Lysosomal dyes and probes are selected not only for signal visibility but also for interpretive clarity, since bright punctate signal can still be misread if workflow conditions are not well controlled. See lysosome staining in cells.
- Cell membrane: Membrane stains are useful when cell outline, boundary definition, or surface-associated visualization is important. See cell membrane staining guide.
- Endoplasmic reticulum: ER staining is often selected for intracellular organization studies or organelle context in multicolor imaging. See endoplasmic reticulum staining.
- Golgi apparatus: Golgi-focused imaging benefits from a workflow that balances probe selection, pattern interpretation, and troubleshooting logic. See Golgi fluorescent staining.
- Cytoskeleton: Cytoskeletal labeling is useful for shape analysis, structural comparison, and spatial interpretation of cell architecture. See cytoskeleton staining.
- Lipid staining: Because lipid labeling depends on the lipophilic behavior of the dye and the hydrophobic nature of the target, stain choice should account for whether the workflow emphasizes general lipid distribution, lipid droplet visualization, or fluorescence-based detection. See lipid staining.
Functional Applications of Cell Staining
Cell staining is often associated with structural visualization, but many workflows also use staining to support function-oriented or process-oriented analysis. In these settings, the objective is not only to show where something is located, but to help distinguish biological states or follow changes over time. These applications require more careful alignment between readout goal and stain logic because the same fluorescent signal cannot always be interpreted as both structure and function without appropriate controls.
- Cell viability staining: Viability-oriented staining is used when the main experimental question is whether cells remain viable, compromised, or non-viable under defined conditions. The value of these assays comes from the ability to distinguish populations clearly rather than simply producing bright fluorescence.
- Cell proliferation tracers: Proliferation tracers are used in workflows where changes associated with cell division or population expansion need to be followed across time or across treatment conditions. The most suitable reagent is usually the one that offers a stable and interpretable relationship to the proliferation question rather than simply the brightest initial signal.
- Cell tracking dyes: Tracking dyes are useful when the goal is to label cells and follow them over time, across compartments, or through broader experimental transitions. These workflows place special weight on signal retention, compatibility with sample handling, and the risk of interpretation drift over extended observation windows.
- Multicolor applications: Functional and structural stains are often combined in multicolor experiments so that one assay can capture more than one dimension of information. This can be highly informative, but it also raises the demands on dye selection, channel planning, brightness balance, and control design. In these settings, panel architecture becomes part of interpretation rather than just image formatting.
Need a practical cell staining strategy for organelle imaging, viability analysis, or multicolor experiments?
We can help align dye selection, staining logic, and channel planning with your sample type, readout priorities, and experimental workflow.
Common Problems in Cell Staining and How to Improve Results
Problems in cell staining are often described in simple visual terms such as weak signal or high background, but the underlying issue is usually broader. A poor result may reflect mismatch between dye and target, inadequate handling conditions, inappropriate imaging timing, or an experimental design that makes the signal hard to interpret. Good troubleshooting therefore asks not only how to make the image look better, but how to make the result more trustworthy.
- Weak or uneven staining: This can arise from low target accessibility, insufficient loading, inconsistent sample preparation, or conditions that do not support stable signal development. The first response should be to review sample health, concentration, incubation, and timing together rather than adjusting only one variable.
- High background fluorescence: Excessive background may result from dye overload, incomplete washing, non-specific retention, or optical interference. In multicolor workflows, background can also be mistaken for target expansion if channel cross-contribution is not considered carefully.
- Poor reproducibility: Reproducibility problems often come from inconsistent sample state, variable imaging settings, or incomplete documentation of staining conditions. A workflow that performs once is not yet a reliable protocol unless it can be repeated under matched conditions.
- Misread signals: The most serious problem is often interpretive. A bright image can still support an unreliable conclusion if staining logic has not been aligned with the biological question. This is especially important in organelle imaging, viability-related workflows, and multicolor experiments.
For a fuller troubleshooting framework organized around causes and solutions, continue to common problems in cell staining and how to fix them.
What is Cell Staining Used For?
Cell staining is a central tool in biomedical and biotechnological research today, which we deploy to characterise the structure, function and pathology of cells. Staining allows for a precise look at various parts of the cell and their internal dynamics, allowing research to advance across multiple disciplines. These are the principal uses of cell staining:
Cell Biology Research
Cell staining is a very common procedure in cell biology to investigate cell structure and function. For instance, by putting particular dyes on the cell membrane, nucleus, mitochondria, endoplasmic reticulum and other parts of the cell, researchers can track cell structure, cell cycle and intracellular transport. Particularly useful for studying the dynamic nature of organelles (mitochondrial fission and fusion, for example) is fluorescent staining of proteins folding and carrying through the endoplasmic reticulum.
Apoptosis and Autophagy Research
This cell staining is used to study apoptosis (programed cell death) and autophagy. With the help of certain dyes (Annexin V-FITC or PI staining), early and late apoptotic cell membrane changes are visible. Also LysoTracker staining is routinely used to study the formation of lysosomes and autophagosomes to reveal how cells, when starved of nutrients or under stress, engage in autophagy. Such research provides an indispensible framework for the pathology of diseases such as cancer and neurodegenerative disorders.
Oncology Research
There are many other uses for cell staining in cancer studies. Using fluorescent labelling of proteins or DNA on tumor cells, scientists can evaluate tumour growth, invasion, and differentiation from normal cells. The molecular substrate of cancer can be identified for instance through Hoechst and DAPI staining, which shows chromatin structure and DNA damage in tumour cells. The cell staining is also used to identify cancer biomarkers which are useful in diagnosing cancer, as they are labelled by specific antigens or proteins.
Cell Differentiation and Regenerative Medicine
Stem cell differentiation, regenerative medicine, they all depend on cell staining. These cells become the specialist cells that they are fluorescently labelled. Neuronal stains (NeuN) follow the differentiation of neuronal cells and muscle stains (a-actinin antibodies) follow the differentiation of muscle cells. And, in tissue engineering and regenerative medicine, the ability to use staining to characterise tissue structure and function guides in vitro construction of living tissue.
Immunology and Infection Research
Cell staining is the science of immune cell recognition and differentiation, as well as immune cell functions in immunology. CD antigen-based staining, for instance, makes multi-layer immune control easily distinguished between various kinds of T cells, B cells and other immune cells. For infectious disease, staining can etch pathogen or host cell antigens to illustrate how viruses, bacteria or other organisms infect and proliferate on host cells. Fluorescent in situ hybridisation (FISH) for instance can identify specific DNA sequences in bacteria infection, and antibody staining can identify the presence of viral proteins in infected cells.
Histology and Pathology Diagnostics
In pathology, staining (H&E staining, PAS staining) are standard tissue lesions diagnosis methods. You can also dye nuclei and cytoplasm in small tissue samples using the H&E technique to observe cell change in size and shape, which can be used by pathologists to diagnose cancer and inflammation. The pathologist increasingly uses fluorescent staining, too. By labelling cancer proteins with antibodies, for instance, we could characterise and molecularly characterise cancers.
Genetics and Molecular Biology Research
Chromosome fluorescence-labelled imaging (FISH, etc) in genetics to depict chromosome evolution and gene loci. FISH can identify exactly which DNA sequences stretch chromosomes to study the relationship between chromosomal breakage and genetic disorders. We also see fluorescent probes made from nucleic acids, which are already ubiquitous in gene expression and regulation studies, and enable us to know what genes do and how they behave molecularly.
How BOC Sciences Supports Cell Staining Workflows?
BOC Sciences supports cell staining workflows by focusing on the real questions researchers usually face before a project becomes reproducible: which stain is appropriate for the target, whether the signal will remain interpretable under the actual workflow, how to reduce avoidable trial-and-error, and how to build a staining strategy that fits imaging, analysis, and downstream decision-making. In practice, customers rarely need "more dyes" in the abstract. They need dyes, probes, and technical support that match their sample type, staining objective, platform conditions, and assay complexity. Our support is therefore organized around common experimental needs, from routine labeling and organelle visualization to multicolor design, mechanism-oriented studies, and customized probe development.
Help with Choosing the Right Stain for the Actual Target
- Support for selecting suitable stains based on what the customer is truly trying to visualize, such as nucleus, membrane, mitochondria, lysosomes, ER, Golgi, cytoskeleton, or broader intracellular organization.
- More practical matching between staining reagents and intended readout, so users can avoid choosing a dye that is bright but poorly aligned with the experimental question.
- Access to commonly used fluorescent dyes and organelle-oriented probes that fit both routine imaging tasks and more target-specific staining workflows.
- Useful starting-point support for teams that know the biological target but need help narrowing the reagent choice according to workflow conditions.
Solutions for Multicolor Imaging and Signal Interpretation Challenges
- Support for customers building multicolor staining panels where channel assignment, brightness balance, and spectral separation matter as much as the individual dye itself.
- Help with selecting dyes that fit around existing reporters, nuclear stains, membrane labels, or organelle probes without creating unnecessary channel conflict.
- More structured evaluation of whether a reagent is suitable for co-staining, confocal imaging, fluorescence microscopy, or broader image-based assay workflows.
- Practical thinking around signal clarity and interpretation, especially when customers need to distinguish real biological pattern from background, bleed-through, or misleading overlap.
Functional Probe Support for Mechanism-Focused Cell Studies
- Probe options relevant to cell viability, cell death, apoptosis-related workflows, cell cycle studies, proliferation analysis, and other function-oriented staining applications.
- Access to functional probes for ROS, pH, Ca2+, and related intracellular response indicators when customers need more than morphology alone.
- Better support for projects where staining is used to compare treatment response, cellular condition, or mechanism-related signal behavior rather than simple endpoint visualization.
- Useful fit for teams developing research workflows that need interpretable fluorescence readouts across pharmacology, response evaluation, and toxicity-related cell analysis.
Technical Guidance for Platform Fit and Experimental Optimization
- Support for matching staining strategies to fluorescence microscopy, confocal imaging, live-cell imaging, and flow cytometry-related workflows.
- Practical consideration of how reagent choice interacts with instrument configuration, acquisition settings, and downstream analysis requirements.
- Useful application support for more complex sample environments, including 3D cell models, organoid-related workflows, and assays with higher imaging or handling demands.
- Experimental optimization guidance intended to help customers reduce troubleshooting burden, improve reproducibility, and move more efficiently from reagent selection to usable staining data.
Do You Need A Consultation?
BOC Sciences integrates cutting-edge fluorescence technologies to accelerate your research, driving next-generation solutions for drug discovery and diagnostics.
Transform Your Studies with Cutting-Edge Fluorescent Products
| Catalog | Name | CAS | Inquiry |
|---|---|---|---|
| A16-0033 | 6-Carboxyfluorescein | 3301-79-9 | Bulk Inquiry |
| A16-0060 | Di-4-ANEPPS | 90134-00-2 | Bulk Inquiry |
| A16-0010 | MitoPY1 | 1041634-69-8 | Bulk Inquiry |
| A16-0005 | MitoMark Green I | 201860-17-5 | Bulk Inquiry |
| A16-0188 | 16:0-DPH PC | 98014-38-1 | Bulk Inquiry |
| A16-0036 | Calcein Blue | 54375-47-2 | Bulk Inquiry |
| A16-0089 | CellHunt CM-DiI | 180854-97-1 | Bulk Inquiry |
| A16-0082 | DiOC16(3) | 161433-32-1 | Bulk Inquiry |
| A16-0156 | Speed DiI | 278173-35-6 | Bulk Inquiry |
| A16-0084 | Speed DiO | 164472-75-7 | Bulk Inquiry |
| A16-0032 | CytoTrace™ Orange CMTMR | 323192-14-9 | Bulk Inquiry |
| A16-0175 | Lucifer Yellow CH Dipotassium Salt | 71206-95-6 | Bulk Inquiry |
| A16-0057 | C10 AV Ceramide | 1263052-40-9 | Bulk Inquiry |
| A16-0051 | C12 NBD Lactosylceramide | 474943-06-1 | Bulk Inquiry |
| A16-0172 | DiBAC4(3) | 70363-83-6 | Bulk Inquiry |
| A16-0031 | DISBAC10 | 169211-45-4 | Bulk Inquiry |
| A16-0083 | Lucifer yellow ethylenediamine | 161578-11-6 | Bulk Inquiry |
High-Performance Fluorescent Tools for Your Research
- Cell Proliferation Tracer Fluorescent Probes Long-term tracking of cell division processes.
- Golgi Fluorescent Probes Targeted probes for Golgi apparatus visualization.
- Nitric Oxide (NO) & Reactive Oxygen Species (ROS) Probes for oxidative stress and signaling detection.
- Ion Fluorescent Probes Indicators for real-time ion concentration imaging.
- Apoptosis Fluorescent Probes Probes detecting programmed cell death events.
- Fluorescent Probes Versatile tools for biomolecular and cellular imaging.
- Other Cell Fluorescent Probes Functional probes for diverse cellular imaging studies.
- Lipid Fluorescent Probes Dyes for lipid droplets and membranes.
Explore More Topics
Online Inquiry