Lysosome Staining in Cells: Choosing Probes and Avoiding Misread Signals
Lysosome staining is widely used in fluorescence-based cell analysis because lysosomes are central to intracellular degradation, vesicle maturation, cargo turnover, and organelle trafficking. In practice, however, lysosome staining is easy to overinterpret. Many researchers expect a bright punctate lysosomal signal to mean they have specifically labeled lysosomes, when in reality a large fraction of commonly used live-cell lysosome probes accumulate because they partition into acidic compartments. That distinction matters. A punctate acidic-vesicle pattern can be highly useful for comparing treatment groups, tracking vesicle redistribution, or adding organelle context to cell imaging workflows, but it does not automatically prove lysosome identity on its own. A useful lysosome staining page therefore needs to do more than introduce LysoTracker dyes. It should explain what these probes really report, where the interpretation boundaries are, why fixation can distort signal meaning, and how to design a staining workflow that reduces false confidence in acidic-compartment readouts.
What Is Lysosome Staining and What It Can Tell You?
Lysosome staining refers to the use of fluorescent dyes or fluorescent probes to visualize lysosome-related signals in cells, most often in live-cell imaging workflows. In practical use, this type of staining is commonly applied to assess the distribution, relative abundance, or acidic behavior of lysosome-associated compartments. However, the interpretation depends strongly on probe mechanism. Many commonly used lysosome stains accumulate because of acidity, so the signal may reflect acidic intracellular compartments rather than lysosome identity alone. For that reason, lysosome staining is useful, but it should be interpreted in relation to the assay goal, staining workflow, and any supporting markers.
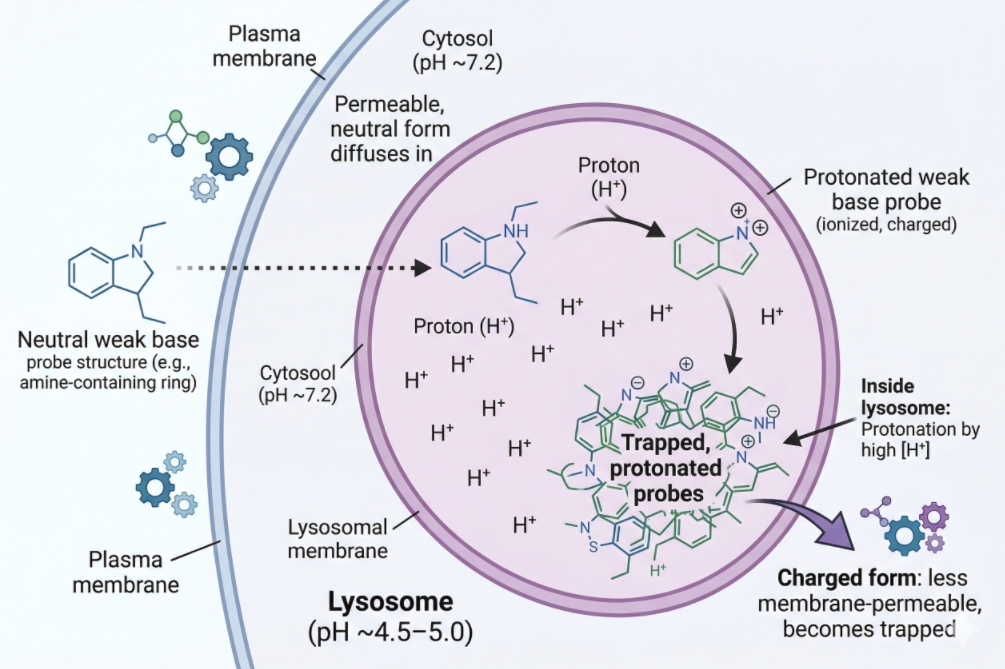 Fig. 1. Many lysosome stains are weak-base probes that accumulate in acidic intracellular compartments in live cells (BOC Sciences Authorized).
Fig. 1. Many lysosome stains are weak-base probes that accumulate in acidic intracellular compartments in live cells (BOC Sciences Authorized).
Definition of Lysosome Staining
At the most basic level, lysosome staining means introducing a fluorescent reagent that helps visualize vesicular structures interpreted as lysosome-related compartments under a fluorescence microscope or comparable imaging platform. In real experiments, this is rarely done only to see lysosomes in a decorative sense. Researchers usually want a practical readout such as punctate vesicle distribution, organelle redistribution after treatment, relative signal change between matched conditions, or spatial relationship between acidic compartments and other cellular structures. This is why lysosome staining often appears in cell staining workflows alongside nuclear dyes, membrane labels, mitochondrial probes, or broader multicolor assay panels. The most common visual output is a punctate intracellular pattern, frequently concentrated in the perinuclear region but sometimes more broadly dispersed depending on cell type, culture condition, treatment exposure, and trafficking state. Researchers often describe this readout using terms such as puncta number, puncta brightness, lysosomal distribution, vesicle clustering, or relative lysosomal intensity. These are legitimate observational outputs. The problem begins when those outputs are treated as if they directly summarize lysosome biology in full. A bright punctate stain does not by itself separate lysosomes from late endosomes or other acidified vesicles, and a weaker stain does not automatically mean lysosomes disappeared.
Information Lysosome Staining Can Provide
When used with the right expectations, lysosome staining can be highly informative. It can reveal whether lysosome-like puncta are evenly distributed or clustered, whether acidic vesicle patterns shift after treatment, whether the cell shows a more compact or more dispersed acidic compartment architecture, and whether lysosome-associated signal changes in parallel with other organelle or morphology readouts. In trafficking-oriented experiments, lysosome staining may help show whether vesicle-rich regions redistribute toward the perinuclear area or toward the cell periphery. In comparative workflows, it can support relative before-versus-after interpretation when imaging settings, probe loading conditions, and handling steps are held constant. Lysosome staining is also frequently integrated into multicolor analysis, where the lysosomal channel adds organelle context rather than functioning as a stand-alone conclusion. For example, a lysosome probe may be combined with a nuclear stain, cytoskeletal label, or viability-associated channel to interpret treatment-induced cellular reorganization more holistically. In those workflows, lysosome staining is useful not because it fully resolves all lysosome biology, but because it adds a compartment-level layer of evidence that can support spatial comparison and hypothesis generation.
Limits of Lysosome Staining for Confirming Lysosome Identity
The most important limitation is simple: acidic accumulation does not equal lysosome-specific recognition. Many popular lysosome dyes, including widely used LysoTracker reagents, are valued because they are convenient for live-cell staining and give a clear punctate signal, but the mechanism is tied to acidic organelle accumulation. That means late endosomal compartments or other acidic vesicles can contribute to the fluorescent pattern. The signal therefore reflects a useful acidic-compartment readout, but not a pure identity marker in the strict sense. Signal intensity is also strongly influenced by lysosomal pH, cell state, treatment condition, incubation timing, dye concentration, and post-staining handling. A treatment that alters vesicle acidification can change the stain even when compartment abundance is unchanged. A stressed or dying cell may show a punctate pattern that looks dramatic but is biologically misleading. If the experiment requires a stronger statement about lysosome identity rather than acidic-vesicle behavior, the stain should be interpreted alongside orthogonal evidence such as an identity-associated marker, a workflow-matched control set, or a separate validation assay. This is the point at which lysosome staining stops being a simple dye step and becomes an interpretation problem.
How Lysosome-Targeting Dyes Work in Live Cells?
To choose a lysosome probe intelligently, it helps to understand the signal mechanism before looking at product names. Most routine live-cell lysosome dyes do not work by highly selective binding to a single lysosomal biomarker. They work because their chemical properties favor accumulation in acidic intracellular organelles. This makes them practical, bright, and easy to use, but it also means the signal is conditional. What you see depends not only on where the compartments are, but on whether the pH gradient is intact, whether the cells remain physiologically stable during staining, and whether imaging happens before the signal redistributes or becomes distorted.
Accumulation in Low-pH Organelles
In live cells, lysosome-targeting dyes such as LysoTracker Blue DND-22, LysoTracker Green DND-26, and LysoTracker Red DND-99 are commonly used because they accumulate preferentially in organelles with acidic luminal environments. Their practical value comes from this pH-associated partitioning behavior. Once inside the cell, the dye enriches in low-pH vesicles and produces a punctate fluorescent pattern that is often interpreted as lysosomal staining. In many routine applications, that is a useful approximation. It allows rapid visualization of acidic vesicle architecture without requiring genetic manipulation or complex immunolabeling steps. The critical nuance is that “low-pH organelles” is not identical to “lysosomes only.” Depending on cell type and biological context, some signal may arise from late endosomes or other acidic compartments that are functionally related but not identical in status. This does not invalidate the stain. It simply changes the wording that should accompany the result. If the biological question is about acidic compartment redistribution, the stain may be well suited. If the question is about confirmed lysosome identity, the dye alone may not be enough.
Signal Retention and Non-Fixable Behavior
One of the most common mistakes in lysosome staining is assuming that a dye performing well in live cells will retain the same meaning after fixation. Many LysoTracker-style reagents are best understood as live-cell probes. Their staining logic depends on the live acidic environment, and once fixation, washing, permeabilization, or prolonged medium exchange is introduced, the original localization meaning may weaken or change. Some signal may remain visible, but visible fluorescence is not the same as preserved interpretive value. This is why LysoTracker fixation questions are so important in assay planning. A researcher may stain live cells, fix them, and still see puncta, yet those puncta may no longer represent the same compartment distribution that existed at the live imaging stage. In other words, signal retention after fixation is not a guarantee of biological fidelity. When a workflow requires fixed-cell imaging, it is safer to test the exact sequence experimentally rather than assuming a live-cell acidic probe can be converted into a robust endpoint identity stain.
Effects of Lysosomal pH Changes on Staining
Because many lysosome probes depend on acidic conditions, any treatment or sample-handling step that shifts lysosomal pH can change the observed readout. This is one of the main reasons lysosome staining needs careful interpretation in perturbation experiments. If a compound, stress condition, starvation protocol, or media change partially neutralizes acidic vesicles, the fluorescence may decrease even though lysosome-associated structures remain present. Conversely, stronger accumulation may sometimes reflect altered pH behavior or loading conditions rather than increased lysosome number. This pH sensitivity is not a flaw; it is part of the probe logic. It becomes a problem only when users treat lysosome staining as a purely static structural marker. In live-cell experiments, the safer interpretation is often that the probe reports the accessible acidic-compartment landscape under those staining conditions. If the experiment is intended to study acidification itself, a pH-sensitive lysosome probe may be more informative. If the goal is to visualize lysosome-like puncta while minimizing functional ambiguity, a more carefully matched workflow or supporting marker may be required.
Choosing between LysoTracker dyes and pH-sensitive lysosome probes?
We can help match acidic-compartment staining logic, channel compatibility, and live-cell workflow needs to your assay goal.
Choosing the Right Probe for Your Experimental Goal
There is no single best lysosome probe for every experiment. The correct choice depends on what the assay is trying to measure, how quickly the sample will be imaged, whether the cells must remain alive, how crowded the spectral panel is, and whether the result needs to say something about acidic compartment behavior or about lysosome identity with greater confidence. A good lysosome probe selection process therefore starts with the readout goal and then moves to spectral fit, workflow stability, and interpretation risk.
When LysoTracker Is a Good Choice?
LysoTracker dyes are a good choice when the experiment needs a practical live-cell readout of acidic vesicle distribution, especially in short imaging windows where convenience, brightness, and organelle contrast matter. LysoTracker Blue DND-22 can be useful when green and red channels are already reserved for other fluorophores. LysoTracker Green DND-26 may be convenient in simpler imaging setups where the green channel is available and detector sensitivity is strong in that region. LysoTracker Red DND-99 is often attractive in multicolor designs because the red channel can reduce conflict with green reporters or other green-emitting cell stains. In these use cases, LysoTracker is valuable because it rapidly reveals acidic puncta and supports comparative live-cell imaging. It is especially useful when the desired output is organelle distribution, relative vesicle pattern change, or inclusion of an acidic-compartment channel in a broader multicolor panel. It is less suitable when the experiment demands hard claims about lysosome identity without additional evidence, or when the workflow depends on post-staining fixation and delayed analysis.
When pH-Sensitive Probes Are More Suitable?
Some experiments are not simply asking where acidic vesicles are located. They are asking whether lysosomal acidification changes under different conditions, whether pH-dependent signal shifts occur after treatment, or whether the assay should emphasize luminal environment rather than localization alone. In those cases, a pH-sensitive lysosome-associated probe may be more suitable than a standard accumulation-based stain. Products such as LysoHunt Yellow HCK-123 may be considered in workflows where acidification-sensitive signal behavior is part of the design logic rather than an unwanted variable. The tradeoff is interpretive complexity. A pH-sensitive probe can be informative, but only if the user explicitly wants that sensitivity. It should not be selected by accident for a morphology-oriented experiment where the real need is stable vesicle visualization. The more the probe depends on pH behavior, the more carefully the experiment must distinguish between compartment presence, compartment acidification, and handling-induced signal change.
How to Choose a Channel-Compatible Probe?
Spectral planning matters in lysosome staining because lysosomal puncta are often bright, compact, and prone to appearing falsely co-localized if neighboring channels are crowded or poorly separated. The best channel is therefore the one that preserves the most important markers in the panel. If the assay already uses a strong green signal for a reporter or cytoskeletal readout, moving lysosome staining to LysoTracker Red DND-99 may reduce channel burden. If red is occupied by another critical marker, LysoTracker Blue DND-22 may be more practical. If the panel is simple and instrument sensitivity in the green region is strong, LysoTracker Green DND-26 can be efficient. Channel choice should also account for autofluorescence, detector performance, filter spacing, and the relative brightness of companion stains. A clean lysosome channel is not just a matter of color preference. It is part of designing a panel that remains interpretable under real acquisition conditions. For broader panel planning logic, researchers often benefit from reviewing how to choose the right fluorescent dye for cell staining before finalizing a lysosome probe.
When an Identity Marker Is Still Needed?
A separate identity-supporting marker is still needed whenever the experiment makes a claim that depends on genuine lysosome assignment rather than acidic-vesicle visualization alone. This includes cases where the manuscript or report needs to distinguish lysosomes from related endolysosomal compartments, where the readout will be used to support mechanistic conclusions, or where treatment effects are expected to alter vesicle pH and trafficking simultaneously. In such scenarios, relying on LysoTracker puncta alone can create a false sense of certainty. The practical standard is not that every lysosome staining experiment must include a second marker, but that interpretation strength should match evidence strength. If the experiment only needs a fast live-cell acidic-compartment readout, LysoTracker may be enough. If it needs to defend lysosome identity more rigorously, orthogonal confirmation becomes far more important.
Live-Cell Workflow for Lysosome Staining
Reliable lysosome staining starts with workflow discipline. Many weak-signal or false-pattern problems are not caused by the reagent itself but by unhealthy cells, mismatched staining medium, overloading, delayed imaging, or inconsistent handling between conditions. Because most routine lysosome probes are used in live cells, the staining workflow should preserve the physiology that makes the acidic compartment signal meaningful in the first place.
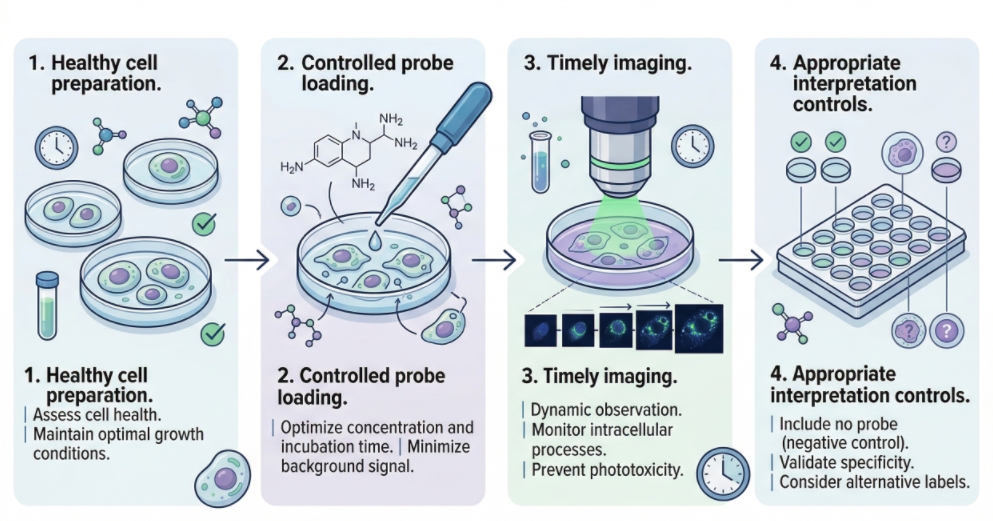 Fig. 2. Live-cell lysosome staining workflow from loading to interpretation (BOC Sciences Authorized).
Fig. 2. Live-cell lysosome staining workflow from loading to interpretation (BOC Sciences Authorized).
1. Cell Preparation Before Staining
Cells should be healthy, appropriately plated, and at a density compatible with clear vesicle visualization. Overcrowded cultures can make puncta appear compressed or merged, while stressed or poorly attached cells often produce distorted acidic-compartment patterns. Before adding a lysosome probe, it is good practice to confirm that the imaging medium, temperature conditions, and handling steps are aligned with the final acquisition setup. A stain optimized in one medium but imaged in another may behave differently, especially if pH buffering or serum conditions change. Cell type also matters. Some cells naturally show dense perinuclear acidic vesicles, whereas others display a broader punctate pattern. That baseline morphology should be established in controls before drawing treatment-related conclusions. In other words, the first question is not whether the puncta look bright, but whether the baseline lysosome-like pattern is stable and believable for that cell model.
2. Probe Concentration and Incubation Conditions
LysoTracker-style probes should be loaded at a concentration that provides clear puncta without saturating the cell with diffuse background. Excessive concentration can raise cytoplasmic haze, overemphasize weak acidic compartments, or make separate vesicles appear artificially fused. Too little probe can leave the image patchy and encourage overprocessing during analysis. Incubation time matters for the same reason. The goal is not maximum brightness; it is an interpretable pattern that remains consistent across matched samples. Because each cell type handles probe loading somewhat differently, pilot optimization is usually more valuable than relying on a generic single-condition assumption. The best test is to compare a modest concentration range under the actual imaging timing you plan to use. A condition that looks ideal immediately after staining may not remain ideal after transfer, washing, or several minutes on the microscope stage.
3. Imaging Timing and Handling After Staining
Live-cell lysosome probes are most defensible when imaged close to the intended staining endpoint under stable environmental conditions. Delays after staining, abrupt medium changes, or extra wash steps can alter distribution, reduce intensity, or introduce stress that changes the very compartments being observed. This is especially important when users are tempted to stain first and decide later whether to image live or fix for downstream analysis. That flexibility often sounds convenient, but it can blur signal meaning. The safer workflow is to decide in advance whether the experiment is a true live-cell imaging assay, a rapid endpoint live readout, or a separate fixed-cell protocol. Once that decision is made, staining, imaging timing, and handling can be optimized as one integrated process rather than as disconnected steps.
4. Controls Needed for Basic Interpretation
Even routine lysosome staining benefits from simple controls. At minimum, unstained cells help define background and autofluorescence. Matched untreated controls establish the baseline acidic-vesicle pattern for the cell type. Workflow-matched treatment controls allow relative comparison under identical acquisition settings. When the biological interpretation depends heavily on acidic-compartment status, an additional perturbation control that is known to affect vesicle acidification can help demonstrate how strongly the signal responds to pH changes. These controls do not make the assay complicated; they make the result more interpretable. Without them, it becomes much easier to confuse differences in staining behavior with differences in lysosome biology.
Multicolor Design Around Lysosome Probes
Lysosome probes are frequently used as one component of a multicolor assay, not as a single-channel experiment. That makes spectral planning and co-localization discipline essential. Bright punctate organelle signals can look deceptively convincing in merged images, and false overlap is common when channels are too close, exposure settings are imbalanced, or one marker is overloaded. A good multicolor lysosome workflow therefore aims to protect interpretability before the images are collected, not after.
Avoiding Channel Overlap in Common Panels
In common three- or four-color panels, one of the easiest mistakes is placing lysosome staining into the most crowded channel simply because that probe is already familiar. Instead, the lysosome channel should be chosen according to channel hierarchy. If the experiment depends on a green biosensor or reporter, using LysoTracker Green DND-26 may create unnecessary bleed-through pressure. In that scenario, LysoTracker Red DND-99 or LysoTracker Blue DND-22 may be more practical. If nuclei are already in blue and the instrument has limited red-channel separation, a green lysosome probe may still be the better compromise. The right answer depends on the whole panel, not the lysosome stain in isolation.
Combining Lysosome Probes with Other Cell Stains
Lysosome probes are often paired with membrane stains, nuclear labels, mitochondrial channels, or readouts related to cell staining quality and morphology. In these settings, the lysosome signal should contribute context rather than dominate the panel. This usually means avoiding unnecessarily bright acquisition settings, keeping puncta resolvable rather than blown out, and ensuring the lysosome channel does not obscure dimmer companion markers. When lysosome staining is combined with assays related to cell health or morphology, it is especially important to remember that stressed cells can change acidic compartment behavior before more obvious damage becomes visible. In some workflows, researchers also pair lysosome staining with viability-oriented readouts to distinguish normal acidic compartment patterning from vesicle behavior in compromised cells. That logic is often more informative than trying to infer cell status from the lysosome channel alone.
Managing Live-Only Probes in Mixed Workflows
Mixed workflows create problems when a live-cell lysosome probe is forced into an endpoint design it was not meant to support. For example, a panel may begin as a live-cell assay but later incorporate fixation, delayed imaging, or post-staining handling to accommodate another marker. Once that happens, the lysosome channel may no longer carry the same meaning it had in the live state. A merged image can still look attractive, but the lysosome interpretation may become weaker. The best solution is to decide early whether the assay is live-first or endpoint-first. If lysosome staining is critical to the question, design the rest of the panel around its live-cell requirements instead of assuming it can be retrofitted into a fixed workflow without consequence.
Reducing False Co-Localization Signals
False co-localization usually comes from one of four sources: spectral bleed-through, overexposure of puncta, cell thickness and projection artifacts, or biological overinterpretation of adjacent vesicles that only appear merged in two-dimensional images. To reduce these problems, each channel should be validated independently, exposure settings should avoid saturation, and co-localization claims should be phrased carefully. “Overlap with acidic compartments” is often more defensible than “lysosome localization” when the lysosome channel itself is based on acidity rather than an identity-specific marker. This wording may sound conservative, but it improves credibility. A more careful description of the data is often stronger than a more aggressive claim built on a weaker signal logic.
Need a cleaner lysosome staining workflow for live-cell or multicolor imaging?
We can help reduce misleading acidic-compartment readouts by aligning probe choice, handling sequence, and panel design with your experiment.
Common Causes of Background and Misinterpretation
Troubleshooting lysosome staining is not only about making the image look cleaner. It is also about preventing misleading conclusions from images that already look acceptable. Some of the most serious lysosome staining problems are interpretive: diffuse background mistaken for vesicle expansion, weak signal mistaken for lysosome loss, or stress-related puncta mistaken for meaningful trafficking biology. Effective troubleshooting therefore has to address both image quality and readout logic.
- Diffuse Background and Overstaining: Diffuse background usually points to one or more of the following: probe concentration is too high, incubation is too long, wash conditions are insufficient, or cell health is poor enough to distort normal compartmental loading. Overstaining can make puncta appear larger, blur compartment boundaries, and increase the temptation to interpret total fluorescence as a biologically meaningful increase. In reality, the image may simply reflect excessive dye. The most practical correction is to reduce loading conditions until puncta remain visible but the cytoplasmic haze drops substantially. A cleaner, slightly less bright lysosome image is usually more interpretable than a brilliant but overloaded one.
- Staining of Other Acidic Compartments: This is the central conceptual problem in lysosome staining. If the probe labels acidic vesicles broadly, then late endosomal or related compartments may contribute to the pattern. In some experiments that is acceptable because the true goal is to monitor the acidic vesicle network as a whole. In other experiments, it becomes a source of overclaiming. The key is to align the language of the conclusion with the actual mechanism of the dye. When there is uncertainty, terms such as acidic compartments, lysosome-associated puncta, or LysoTracker-positive vesicles can be more accurate than declaring definitive lysosome identity.
- Signal Distortion After Fixation or Media Changes: Fixation, temperature drift, imaging-medium exchange, and prolonged delay after staining can all distort lysosome probe behavior. The resulting image may still show puncta, but those puncta may no longer match the live acidic-compartment pattern that the probe originally reported. This is especially relevant in workflows where users stain in one buffer, wash into another, then image later under non-matched conditions. Small changes can alter organelle behavior enough to complicate interpretation. When signal meaning matters, the exact handling sequence should be treated as part of the assay design, not as an afterthought. This is one reason why disciplined cell staining protocol planning reduces troubleshooting later.
- Misleading Patterns in Stressed or Dying Cells: Stressed cells often show abnormal vesicle patterns, altered pH regulation, condensed morphology, or fragmented intracellular organization that can make lysosome staining look highly dramatic. These images are easy to overinterpret because they contain bright puncta and obvious structural change. Yet they may reflect general cell compromise rather than a specific lysosome-centered biological event. When treatment conditions are harsh, a companion readout related to morphology or viability is often necessary to avoid mistaking a dying-cell phenotype for a meaningful lysosome trafficking result. This is where general cell staining troubleshooting logic becomes important. If the lysosome image looks striking but the cells themselves appear unhealthy, the first task is not to explain the biology in depth. It is to determine whether the assay still supports a valid biological interpretation.
Best Practices for Reliable Lysosome Readouts
The most reliable lysosome staining workflows are built by matching probe logic to readout goal, validating the live-cell sequence under real imaging conditions, and reporting conclusions at the strength supported by the data. This approach is more valuable than simply selecting the brightest dye. A robust lysosome assay is one where users know what the stain is showing, what it is not proving, and when additional evidence becomes necessary.
- Matching the Probe to the Readout Goal: Before selecting a product, define whether the experiment needs one of the following: a practical live-cell acidic compartment stain, a pH-sensitive readout, a multicolor-compatible vesicle channel, or a stronger lysosome identity workflow. Once the goal is explicit, selection becomes easier. LysoTracker Green DND-26, LysoTracker Red DND-99, and LysoTracker Blue DND-22 are most useful when fast live-cell acidic-compartment visualization is the priority. LysoHunt Yellow HCK-123 may be more informative when pH-linked interpretation is the intended readout. The wrong probe is often not chemically poor; it is simply mismatched to the experiment.
- Using Supporting Markers When Interpretation Matters: Additional markers or orthogonal validation become more important as the interpretation becomes more specific. If the result will be described as lysosome redistribution, lysosome co-localization, or lysosome abundance change in a strong mechanistic sense, supporting evidence should increase accordingly. This does not mean every assay needs a complex validation tree. It means the wording of the conclusion should be proportionate to the specificity of the evidence.
- Deciding When to Repeat or Escalate the Assay: A repeat or escalation is warranted when signal changes are large but cell health is questionable, when puncta appear convincing only under aggressive image processing, when fixation or handling has altered the workflow substantially, or when the biological conclusion depends on separating lysosomes from other acidic compartments. In those cases, the stain may still be useful as a first-pass readout, but not as the only layer of evidence. Repeating the assay with cleaner controls, improved channel allocation, or an added identity-supporting step is often more efficient than defending an ambiguous image after the fact.
- Reporting Results in a More Defensible Way: The strongest reporting language is often the most precise. Instead of saying that a LysoTracker signal definitively labels lysosomes in all circumstances, it is usually safer to state that the probe visualized acidic vesicular compartments consistent with lysosome-associated staining under the tested live-cell conditions. Instead of claiming absolute lysosome increase from intensity alone, it is often better to describe a relative increase in LysoTracker-positive puncta or acidic-compartment fluorescence under matched conditions. These distinctions may seem subtle, but they greatly improve the credibility of the result.
How BOC Sciences Supports Lysosome Staining Workflows?
BOC Sciences supports lysosome staining workflows through integrated capabilities in fluorescent product supply, custom probe development, staining strategy optimization, and technical service support. Rather than treating lysosome staining as a simple puncta-labeling step, we help researchers align probe mechanism, acidic-compartment selectivity, spectral placement, and imaging conditions with the actual purpose of the experiment. This is especially important in lysosome-related imaging, where signal behavior may depend on compartment acidification, probe accumulation, live-cell handling, and multicolor panel design. Our support is therefore structured to cover both routine lysosome visualization needs and more specialized fluorescence assay development requirements.
Lysosome Fluorescent Dye and Probe Supply
- Access to a range of lysosome-associated fluorescent dyes and probes suitable for acidic-compartment visualization, puncta imaging, and routine lysosome-related fluorescence workflows.
- Support for selecting lysosome probes across different spectral regions, including blue, green, yellow, red, and longer-wavelength fluorescence options for more flexible multicolor assay design.
- Availability of probe types suited to different readout needs, including accumulation-based lysosome stains, pH-responsive fluorescent probes, and related reagents for compartment-focused imaging.
- Product support spanning both standard lysosome-labeling reagents and more specialized fluorescent materials relevant to organelle interaction studies and vesicle-oriented analysis.
Customized Lysosome Probe Development
- Custom design of lysosome-targeted fluorescent probes according to specific research goals, imaging platforms, and desired compartment-labeling behavior.
- Structural modification support for fluorescent dyes, including adjustment of pKa-related behavior, hydrophobicity, charge, and targeting features to improve lysosome-associated localization and signal performance.
- Development support for functional probe systems when lysosome targeting needs to be combined with responsive fluorescence logic, such as pH-sensitive, enzyme-related, ion-associated, or oxidative-state-linked readouts.
- One-stop development services covering molecular design, synthesis, modification, purification, and characterization for customized lysosome fluorescent probes.
Lysosome Staining Workflow Optimization and Application Support
- Practical guidance for optimizing lysosome staining conditions, including dye concentration, incubation time, temperature, medium choice, washing strategy, and imaging timing.
- Application support for different experimental systems, including live cells, fixed-cell endpoint workflows, suspension cells, adherent cells, and broader image-based assay formats where lysosome channels must remain interpretable.
- Suggestions for multicolor staining strategies involving lysosomes together with nuclei, mitochondria, endoplasmic reticulum, membranes, or other organelle markers in broader fluorescence panels.
- Troubleshooting assistance for common lysosome staining challenges, including diffuse background, overstaining, weak puncta contrast, handling-induced signal drift, or uncertainty in acidic-compartment interpretation.
Quality Control and Technical Service Support
- Supply of high-purity lysosome-related fluorescent products supported by strict quality control processes, with analytical methods such as HPLC, NMR, and MS used where applicable.
- Attention to batch-to-batch consistency and product stability to support reproducibility across repeated lysosome staining and imaging experiments.
- Availability of complete technical documentation, including COA, MSDS, and product application information, to support evaluation, workflow planning, and implementation.
- Access to professional technical support for both pre-sale consultation and post-sale troubleshooting, helping researchers address issues from probe selection to lysosome imaging optimization.
Do You Need A Consultation?
BOC Sciences integrates cutting-edge fluorescence technologies to accelerate your research, driving next-generation solutions for drug discovery and diagnostics.
Transform Your Studies with Cutting-Edge Fluorescent Products
| Catalog | Name | CAS | Inquiry |
|---|---|---|---|
| A16-0183 | NBD-C6-HPC | 81005-34-7 | Bulk Inquiry |
| A16-0072 | 5-Dodecanoylaminofluorescein | 107827-77-0 | Bulk Inquiry |
| A16-0090 | 3-NBD-C12 Cholesterol | 186833-02-3 | Bulk Inquiry |
| A16-0101 | LysoHunt Yellow HCK-123 | N/A | Bulk Inquiry |
| A16-0094 | LysoTracker Blue DND-22 | N/A | Bulk Inquiry |
| A16-0131 | LysoTracker Green DND-26 | N/A | Bulk Inquiry |
| A16-0137 | PFB-FDGlu | N/A | Bulk Inquiry |
| A16-0191 | MHI-148 | 172971-76-5 | Bulk Inquiry |
| A16-0206 | C-Laurdan | 959839-06-6 | Bulk Inquiry |
| A16-0194 | LysoTracker Red DND-99 | 231946-72-8 | Bulk Inquiry |
High-Performance Fluorescent Tools for Your Research
- pH Indicators Fluorescent sensors for intracellular pH monitoring.
- Mitochondrial Fluorescent Probes Targeted dyes for mitochondrial structure and function.
- Cell membrane Fluorescent Probes Surface-labeling dyes for membrane dynamics studies.
- Ion Fluorescent Probes Indicators for real-time ion concentration imaging.
- Calcium, Chloride and Other indicators Fluorescent indicators for intracellular ion flux monitoring.
- Lysosomal Fluorescent Probes Acidic organelle markers for lysosome tracking.
- Golgi Fluorescent Probes Targeted probes for Golgi apparatus visualization.
- Endoplasmic Reticulum Fluorescent Probes ER-targeted dyes for organelle structure analysis.
Explore More Topics
Online Inquiry