Organelle-Specific Cell Staining: Mitochondria, Nucleus, Membrane and Beyond
Organelle cell staining is not just a more detailed version of general cell staining. For many researchers, it is the point where a fluorescence experiment becomes genuinely informative. When the question is no longer "Are the cells there?" but rather "What is happening inside the cells?", broad whole-cell contrast quickly becomes insufficient. Researchers need to know whether mitochondria are redistributed, whether lysosomes are expanded or clustered, whether the plasma membrane remains well defined, whether the nucleus can serve as a reliable reference layer, and whether lipid-associated structures are changing in ways that affect interpretation. In these situations, organelle-specific staining is not a visual preference. It is a decision about analytical precision. This is also why organelle staining is one of the most misunderstood areas in fluorescence workflow design. A stain that appears bright and visually attractive can still fail the real experimental need if it does not reflect the correct compartment, if its signal depends on a functional property that changes during sample handling, or if it cannot be combined reliably with other dyes in a multicolor experiment. Many staining failures happen because researchers choose an organelle stain based on familiarity, color, or a general label claim without fully checking how the stain enters cells, what drives retention, whether it remains interpretable after fixation, and how it behaves alongside other channels. The result may be a strong image but a weak conclusion.
What Is Organelle Cell Staining?
Organelle-specific cell staining is the use of fluorescent dyes, fluorescent probes, or labeled reagents that localize preferentially to a defined intracellular compartment so that the compartment can be visualized, compared, or analyzed in the context of the whole cell. That definition may sound straightforward, but in real experimental design it carries several important implications. First, the signal is only useful if it corresponds closely enough to the intended organelle to support biological interpretation. Second, specificity is not a binary marketing claim but a practical property shaped by sample type, staining conditions, live-versus-fixed workflow, washing strength, and acquisition settings. Third, some organelle stains act mainly as localization tools, while others report on the environment or functional state of the compartment they label. A researcher selecting an organelle stain therefore needs more than a category name. They need to understand what the signal is supposed to mean under their actual workflow conditions.
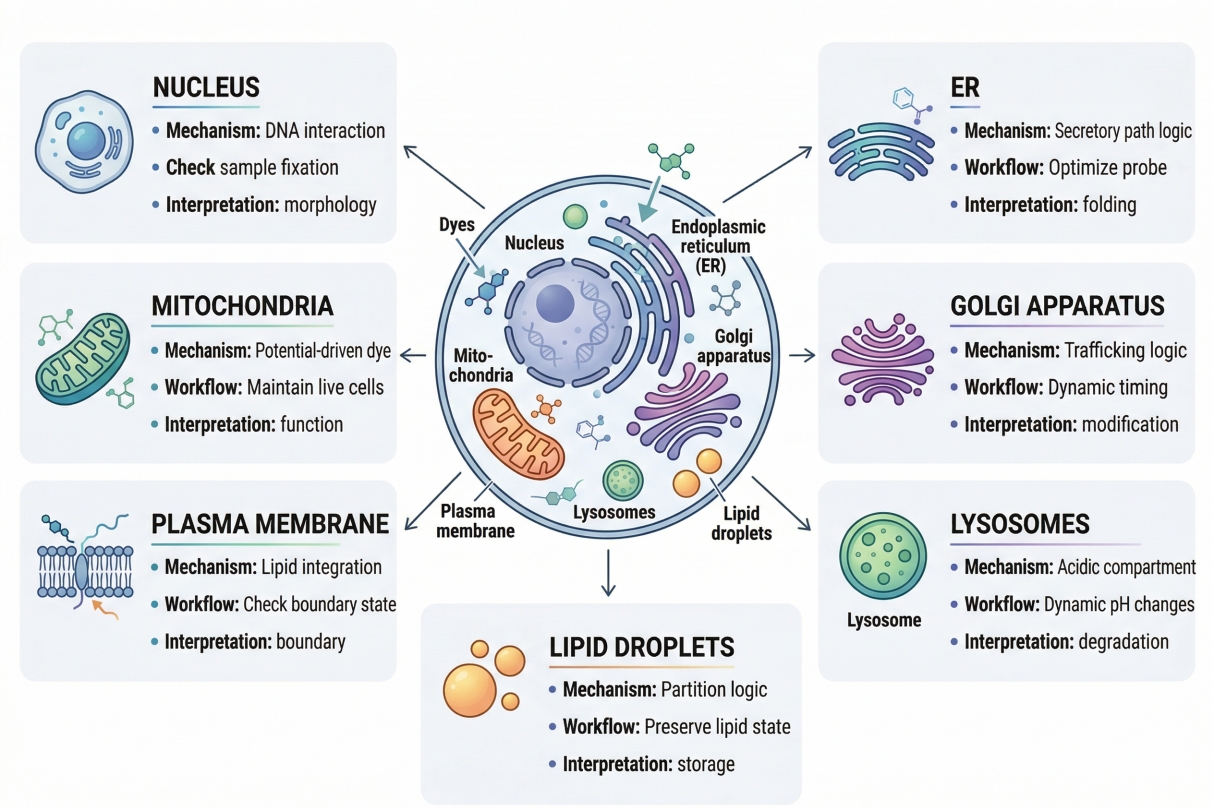 Fig. 1. Different organelles require different staining mechanisms, workflow conditions, and interpretation logic (BOC Sciences Authorized).
Fig. 1. Different organelles require different staining mechanisms, workflow conditions, and interpretation logic (BOC Sciences Authorized).
In practical terms, organelle-specific staining means narrowing the fluorescence readout from the whole cell to a particular intracellular structure such as the nucleus, mitochondria, plasma membrane, lysosomes, ER, Golgi apparatus, or lipid-associated compartments. That narrowing is what makes the method valuable. Instead of collecting broad intracellular fluorescence and trying to infer which region matters, the researcher deliberately chooses a probe whose chemical behavior increases signal within the target compartment more than elsewhere. Depending on the probe, this may occur through nucleic acid binding, membrane partitioning, acidic-compartment accumulation, electrochemical preference, lipid interaction, or another mechanism. The key point is that organelle-specific staining is an analytical method for targeting intracellular information, not simply a way to decorate the cell with color.
How Organelle Staining Differs from General Cell Staining?
General cell staining is often used to show broad morphology, confirm the presence of cells, or provide a whole-cell or reference-level visual layer. Organelle staining has a higher interpretive burden because the experimental claim is more specific. If a whole-cell stain is somewhat diffuse, it may still be useful for rough morphology. If an organelle stain is diffuse, its value may collapse because the experiment depends on knowing where the signal is within the cell. This difference is highly relevant to researchers who need clear localization rather than general contrast. A researcher studying organelle interaction, intracellular trafficking, or treatment-induced redistribution needs specificity at the compartment level, not just fluorescence somewhere in the cytoplasm. That is why organelle staining choices should be made more carefully than many entry-level protocols suggest.
Why Organelle Specificity Matters in Fluorescence-Based Cell Analysis?
Organelle specificity matters because many cellular responses are spatial before they become grossly visible. A treatment may not kill cells outright, but it may fragment mitochondria, enlarge lysosome-associated structures, alter membrane organization, shift the Golgi pattern, or change the relationship between lipid-rich domains and the rest of the cell. These are precisely the kinds of changes that researchers often need to detect in early screening, mechanistic follow-up, or image-based comparison studies. Without organelle specificity, those changes are diluted into broad intracellular signal and become difficult to distinguish from background or subjective interpretation. With appropriate organelle staining, the same sample can provide structurally anchored, compartment-level information that is far more actionable in microscopy, fluorescence imaging, and multicolor assay design.
Why Different Organelles Require Different Staining Strategies?
A common researcher assumption is that once a fluorescent dye works for one organelle, selecting a second or third organelle stain is mainly a matter of choosing another color. In reality, different organelles require different staining strategies because they are chemically and structurally different environments. The nucleus is not labeled for the same reason as a lysosome. A plasma membrane stain does not behave like a mitochondria-oriented probe. An ER network cannot be interpreted in the same way as a compact Golgi-associated signal. This means organelle staining is not a "one logic, many colors" exercise. It is a set of related but distinct strategies whose success depends on understanding why the target compartment is stainable in the first place.
Differences in Membrane Composition, Permeability, and Intracellular Accessibility
Different organelles differ in membrane composition, internal environment, accessibility from the extracellular space or cytosol, and the extent to which they can retain a probe during washing or processing. These differences matter directly to researchers because they determine whether a stain will localize rapidly, whether it will need living cells to behave correctly, and whether the signal will remain interpretable after fixation. For example, a nucleus-oriented stain often relies on DNA-rich access and strong nucleic acid interaction, whereas plasma membrane dyes frequently depend on surface membrane association or lipid partitioning. Lysosome-associated probes are often shaped by acidic compartment behavior, while lipid-oriented probes may reflect membrane order or lipid-rich regions rather than a classic membrane-bounded organelle. A stain that ignores these differences may still fluoresce, but it may not answer the organelle-specific question the experiment was designed to solve.
Structural Staining vs Environment-Dependent Staining Mechanisms
Another major reason organelles need different strategies is that not all organelle stains are purely structural. Some are closer to reference labels whose main value is where they localize. Others are environment-dependent and therefore reflect not only the presence of the organelle but also a compartment property such as membrane potential, acidity, lipid packing, or another physiochemical condition. This distinction matters enormously in researcher workflows where the goal is to compare treated and untreated samples. If the probe is function-sensitive, a change in signal may reflect altered compartment state rather than more or less of the compartment itself. Researchers need that distinction clearly because it changes how results should be interpreted, how controls should be designed, and whether the same stain can be trusted in live-cell versus fixed-cell analysis.
Why Live-Cell Compatibility and Fixation Behavior Vary Between Organelle Stains?
Live-cell compatibility varies between organelle stains because the biological drivers of staining are not universal. Some organelle probes behave well only while cells remain viable and compartment conditions are intact. Others tolerate fixation more effectively because their localization depends less on transient physiology and more on stable structural association. From a researcher-needs perspective, this is often where staining workflows fail: a stain is selected because it works beautifully in a live-cell product image, but the actual project requires fixation, permeabilization, delayed imaging, or combination with downstream labels. Once the workflow changes, the signal may weaken, redistribute, or lose biological meaning. That is why organelle stain selection should always be tied to the full workflow, including whether the sample remains live, whether it will be fixed, and whether the signal must coexist with later processing steps.
Need help choosing the right organelle staining workflow for imaging or assay development?
We can help match stain specificity, signal performance, and channel placement to your cell type, experimental design, and microscopy setup.
Nuclear Staining: DNA-Associated Labeling and Reference Signal Design
Nuclear staining is often the first organelle-level channel established in fluorescence workflows because it provides both a technically stable reference layer and a biologically useful readout. In research settings across pharma, biotech, new drug discovery, and CDMO laboratories, nuclear staining is frequently used not only to visualize nuclei, but also to support cell counting, segmentation, normalization across treatment groups, and image-based comparison of morphology or intracellular distribution. For this reason, nuclear staining should be planned as a functional part of assay architecture rather than as a simple counterstain added at the end.
- Technical Basis of Nuclear Staining: Most nuclear staining strategies rely on strong interaction with nucleic-acid-rich structures, especially DNA, which creates high contrast between the nucleus and surrounding cytoplasmic space. From a technical perspective, this makes nuclear staining one of the most robust fluorescence channels when the goal is to define cell number, identify nuclei for segmentation, or create a stable positional anchor for additional organelle or marker channels. However, the apparent simplicity of nuclear staining can be misleading. Signal behavior still depends on permeability, fixation compatibility, spectral channel assignment, and the extent to which the staining conditions preserve clear nuclear boundaries without generating excessive background or oversaturation.
- Application Value in Imaging, Screening, and Comparative Analysis: In applied research workflows, nuclear staining is widely used in high-content imaging, phenotypic screening, intracellular localization analysis, and treatment-response studies because it enables normalization of fluorescence data to cell number and supports reproducible field-to-field comparison. In multicolor experiments, the nuclear channel often serves as a structural anchor that helps interpret whether other signals are perinuclear, cytoplasmic, membrane-associated, or organelle-restricted. This is particularly useful in compound screening and mechanism-of-action studies where researchers need consistent spatial reference points across many images, conditions, or time points rather than isolated visual snapshots.
- Key Risks and Design Considerations: The main technical risks in nuclear staining are overloading, spectral crowding, and misinterpretation of abnormal nuclear morphology. Excessively intense nuclear signal can mask internal nuclear detail and complicate segmentation, while inappropriate channel placement can interfere with weaker but biologically critical fluorophores in multicolor designs. In stressed or treatment-exposed samples, nuclear shape may also change substantially, which means the nuclear channel should not be treated as automatically neutral. For robust assay design, nuclear staining should be selected according to workflow type, image analysis needs, and panel structure rather than used as a default channel without optimization.
Mitochondrial Staining: Localization, Functional Sensitivity, and Signal Interpretation
Mitochondrial staining is among the most information-rich organelle staining strategies because it can reveal both intracellular organization and biologically relevant changes in organelle state. In translational and discovery-oriented research, mitochondrial channels are often incorporated into assays designed to compare compound response, assess intracellular stress, monitor organelle redistribution, or support multiparameter image analysis. The value of mitochondrial staining is therefore not limited to making the organelle visible. Its real utility lies in how well the selected probe matches the intended interpretation.
- Technical Basis of Mitochondrial Staining: Mitochondrial stains generally fall into two broad technical groups: probes used primarily for localization and probes whose accumulation or fluorescence is influenced by mitochondrial physiology. This distinction is critical because a mitochondrial signal does not always represent organelle abundance alone. Depending on the probe, signal intensity may also reflect membrane-associated properties or other conditions that affect probe retention. From a technical standpoint, researchers therefore need to understand whether the selected stain is mainly a structural marker, a function-sensitive probe, or something in between. Without that distinction, intensity changes can easily be overinterpreted.
- Application Value in Drug Discovery and Cellular Mechanism Studies: In pharma, biotech, and CDMO environments, mitochondrial staining is especially valuable in assays that examine treatment-induced redistribution, organelle morphology changes, intracellular stress response, or relationships between mitochondria and other compartments such as nuclei, lysosomes, or membranes. It is commonly used in live-cell imaging to preserve dynamic information and in image-based screening to compare intracellular phenotype changes across compounds or conditions. A well-designed mitochondrial channel can therefore support both descriptive imaging and mechanistic interpretation, provided the staining logic remains aligned with the biological question.
- Key Risks and Design Considerations: The major risks in mitochondrial staining arise when function-sensitive signal is interpreted as if it were purely structural, or when live-cell-derived staining logic is carried into fixed-cell analysis without validation. Mitochondrial fluorescence may change because of altered organelle condition, not simply because the organelle has redistributed or changed in abundance. In addition, fixation and post-processing can change retention behavior and reduce interpretability. For robust experimental design, mitochondrial probes should be selected according to whether the assay requires live-cell sensitivity, endpoint structural visualization, or compatibility with broader multicolor panel architecture.
Plasma Membrane Staining: Boundary Definition, Cell Morphology, and Surface-Oriented Imaging
Plasma membrane staining is technically valuable because it creates a clear boundary layer that supports cell segmentation, morphology assessment, and interpretation of how intracellular signals relate to the cell surface. In research workflows relevant to therapeutic screening, cell-based assay development, and imaging-driven mechanism studies, membrane staining is often used to improve single-cell definition in crowded fields, distinguish surface-associated from intracellular localization, and stabilize multicolor image interpretation. Its usefulness is therefore closely tied to image quality and assay structure rather than to membrane labeling alone.
- Technical Basis of Membrane Staining: Membrane stains are usually designed to associate with the plasma membrane through surface-oriented or membrane-partitioning behavior, producing fluorescence that outlines the cell boundary more clearly than general cytoplasmic contrast. Technically, the challenge is not only obtaining signal, but preserving a signal pattern that remains sufficiently surface-restricted to be analytically useful. If the stain partitions too broadly into internal membrane systems, becomes internalized too rapidly, or is used under conditions that favor diffuse accumulation, the channel may remain bright while losing value as a true boundary marker.
- Application Value in Morphology Analysis and Multicolor Imaging: In application-driven settings, membrane staining is particularly useful for morphology-sensitive workflows, cell-shape comparison, segmentation-assisted image analysis, and multicolor panels where a reliable cell outline improves interpretation of nuclear, organelle, or viability-related channels. In high-content imaging and phenotypic screening, a well-resolved membrane channel can improve cell identification and reduce ambiguity when multiple cells occupy the same field. It is also valuable when the biological question concerns whether a signal is confined to the surface, redistributed internally, or associated with changes in cell boundary organization after treatment.
- Key Risks and Design Considerations: The most common design problem in membrane staining is assuming that any lipophilic or membrane-associated fluorescence is equivalent to high-quality plasma membrane labeling. In practice, diffuse staining, internalization, excessive loading, and post-fix redistribution can all compromise the usefulness of the channel. For robust workflows, membrane stains should be chosen with attention to live versus endpoint imaging needs, compatibility with other channels, and whether the signal must function primarily as a morphology reference, a dynamic membrane readout, or a supporting layer in multicolor analysis.
Lysosome Staining: Acidic Compartment Labeling and Interpretation Challenges
Lysosome staining is frequently incorporated into organelle-oriented imaging workflows because acidic compartments are relevant to intracellular trafficking, stress response, organelle interaction, and treatment-related phenotype analysis. In screening and mechanistic studies, lysosome-associated fluorescence can provide a strong punctate or compartmental signal that adds substantial biological depth to a panel. However, lysosome staining also requires more careful interpretation than many structure-oriented channels because the readout is often closely tied to compartment chemistry.
- Technical Basis of Lysosome Staining: Many lysosome-targeted stains depend on acidic compartment behavior for accumulation or signal enrichment, which means the fluorescence pattern reflects more than simple organelle presence. Technically, this is useful because it can create strong contrast within acidic structures, but it also means that the signal can shift when compartment chemistry changes. Lysosome-associated fluorescence therefore needs to be understood as condition-dependent in many workflows. A change in staining pattern may reflect altered acidic behavior, altered compartment organization, or both.
- Application Value in Trafficking, Stress, and Organelle Interaction Studies: Lysosome staining is especially valuable in research programs where intracellular trafficking, vesicle-related organization, or treatment-dependent compartment behavior must be compared across conditions. It is commonly used in multicolor imaging to define lysosome position relative to nuclei, membranes, or mitochondria and to assess whether compounds or perturbations alter compartment distribution. In drug discovery and cell-based assay contexts, lysosome channels can add important interpretive context to phenotypic imaging by showing whether intracellular changes are associated with altered acidic compartment patterning rather than only with broad morphology differences.
- Key Risks and Design Considerations: The main risk in lysosome staining is overinterpreting the signal as purely structural when it is partly environment-dependent. Sample stress, media changes, incubation conditions, and post-staining processing can all influence signal behavior. As a result, a lysosome-oriented channel may appear to change significantly even when the underlying biology is more subtle than the fluorescence suggests. For robust assay design, lysosome stains should be validated under the exact sample handling conditions used in the experiment and should be interpreted in relation to control conditions that clarify whether signal differences reflect compartment abundance, compartment state, or technical workflow effects.
Endoplasmic Reticulum Staining: Intracellular Network Visualization and Workflow Considerations
Endoplasmic reticulum staining is useful in workflows where researchers need to visualize intracellular architecture more deeply than nucleus-plus-cytoplasm channels allow. Because the ER forms an extended reticular network, it provides a different kind of information from compact organelles such as the Golgi or punctate lysosome-associated structures. In research settings focused on intracellular organization, multiorganelle relationships, or treatment-driven structural phenotype analysis, ER staining can be a valuable channel when its network character is preserved clearly.
- Technical Basis of ER Staining: ER-oriented staining is technically distinct because the target is not a compact, isolated compartment but a broad intracellular membrane network. The signal must therefore be judged by pattern quality as much as by brightness. A technically useful ER channel should reveal a coherent reticular organization rather than merely increasing generalized intracellular fluorescence. This means ER staining is especially sensitive to stain choice, loading behavior, background control, and how the workflow affects intracellular membrane architecture during imaging or post-processing.
- Application Value in Intracellular Organization Studies: In applications such as morphology-sensitive imaging, organelle relationship analysis, and complex phenotypic screening, ER staining helps define internal architecture that cannot be resolved through nuclear or membrane channels alone. It can provide context for interpreting how intracellular structures are distributed relative to one another and whether a treatment disrupts or reorganizes the internal network of the cell. For researchers in drug discovery or biologics development, this can be useful when cell state changes are reflected more in internal organization than in gross morphology.
- Key Risks and Design Considerations: The main interpretation problem in ER staining is confusing broad intracellular fluorescence with authentic network labeling. Because the ER occupies a large fraction of the cell interior, poor specificity or excessive staining can produce patterns that look informative while lacking structural precision. In addition, fixation and other processing steps can change the apparent continuity or sharpness of the network. For robust results, ER stains should be selected and optimized with attention to whether the workflow requires live visualization, endpoint comparison, or compatibility with multiple additional organelle channels.
Golgi Staining: Compact Organelle Labeling and Spatial Organization Analysis
Golgi staining is particularly valuable in imaging workflows where a compact intracellular reference is needed to evaluate spatial organization and treatment-related redistribution within the cell. Compared with extended networks such as the ER, the Golgi often provides a more localized signal that can be useful for comparing intracellular structure across experimental conditions. This makes it attractive in image-based mechanism studies and organelle relationship analysis, provided the staining pattern remains sufficiently coherent to support interpretation.
- Technical Basis of Golgi Staining: Technically, a useful Golgi stain should generate a localized intracellular pattern that can be distinguished from diffuse cytoplasmic fluorescence. Because the Golgi is spatially compact relative to broader membrane systems, the success of the stain depends heavily on localization fidelity rather than raw intensity. This makes Golgi labeling sensitive to processing effects, intracellular redistribution, and background contributions that can blur the boundary between a true Golgi-associated pattern and general intracellular fluorescence.
- Application Value in Spatial Organization and Comparative Imaging: In application terms, Golgi staining is valuable when researchers need to compare how intracellular organization changes across treatment groups, cell states, or assay time points. It can function as a spatially informative internal reference in microscopy and can help define whether intracellular organization is preserved, shifted, or disrupted. In research environments focused on compound response or phenotypic comparison, this type of compact organelle channel can add interpretive clarity that broader cytoplasmic contrast cannot provide.
- Key Risks and Design Considerations: The main risk in Golgi staining is overcalling specificity based on compact appearance alone. Perinuclear or localized intracellular fluorescence may appear Golgi-like without being sufficiently target-restricted for confident interpretation. Processing can also alter apparent position or compactness, which is especially important in comparative workflows. For robust use, Golgi staining should be optimized with attention to localization quality, consistency across conditions, and whether the channel is being used as a descriptive marker, a comparative reference, or part of a broader multiorganelle panel.
Lipid-Associated Compartment Staining: Membrane-Rich Domains, Droplets, and Beyond
Lipid-associated compartment staining is especially relevant in workflows where membrane-rich structures, lipid-environment changes, or lipid-associated intracellular organization contribute to the experimental question. This area is broader and more technically variable than many classical organelle channels because lipid-oriented probes often respond to membrane interaction, membrane order, or lipid-rich environments rather than labeling one single bounded organelle. As a result, these stains can be highly informative, but only when the biological question and the signal mechanism are aligned clearly.
- Technical Basis of Lipid-Associated Staining: Lipid-associated stains typically work through interaction with membrane-rich or lipid-associated environments, which means their fluorescence behavior can depend on partitioning, local membrane properties, or related physicochemical conditions. Technically, this makes them different from highly discrete organelle labels. The key issue is not simply whether fluorescence appears in lipid-relevant regions, but whether the observed pattern is specific enough to support the type of compartment-level interpretation the experiment requires. In some workflows, broad membrane-associated contrast is sufficient. In others, more discrete lipid-related structures or domains must be resolved.
- Application Value in Membrane Biology and Phenotypic Imaging: In application-driven settings, lipid-associated staining can support studies of membrane organization, lipid-rich compartment behavior, phenotypic response, and multiorganelle context where membrane-related changes are part of the biological mechanism being investigated. These channels can add value in phenotypic screening and microscopy-based analysis when compound treatment, formulation exposure, or intracellular stress is expected to influence membrane-associated organization. They are particularly useful when a conventional nucleus–membrane–organelle panel would otherwise miss biologically important lipid-environment information.
- Key Risks and Design Considerations: The most important design risk is treating all membrane-associated fluorescence as if it were highly specific for one lipid-rich compartment. In practice, lipid-oriented signal may reflect broad membrane interaction rather than a single defined organelle, and changes in intensity may arise from altered partitioning rather than from simple increases or decreases in compartment abundance. For robust interpretation, lipid-associated stains should be chosen with careful attention to what the assay actually needs to reveal, whether the workflow is structural or state-sensitive, and how the signal will be distinguished from broader membrane-associated background.
Cytoskeleton Staining: Structural Organization, Morphology Readouts, and Imaging
Cytoskeleton staining is especially valuable in imaging workflows where cell shape, intracellular architecture, and structural redistribution need to be interpreted under controlled experimental conditions. Because the cytoskeleton spans the cell through organized filament systems rather than through a compact organelle-like compartment, a useful stain should do more than generate bright intracellular contrast. It should help reveal whether filament organization remains coherent, whether structural pattern changes after treatment, and whether the observed signal supports a meaningful interpretation of cell organization rather than a generic increase in intracellular fluorescence.
- Technical Basis of Cytoskeleton Staining: Technically, cytoskeleton staining is intended to visualize filament-associated structures such as actin-rich networks or microtubule-based intracellular architecture in a way that preserves interpretable structural pattern. Because these elements extend across the cell and are closely tied to morphology, staining quality depends heavily on resolution, background control, and preservation of filament continuity rather than on signal intensity alone. A strong signal is only useful when it remains spatially organized enough to distinguish real structural features from diffuse intracellular labeling.
- Application Value in Morphology and Comparative Imaging: In practical imaging studies, cytoskeleton staining is highly valuable when researchers need to compare cell spreading, filament redistribution, edge structure, intracellular tension-related patterning, or treatment-associated changes in overall cellular organization. It can provide an informative structural layer in assays focused on adhesion, migration, morphology, or multicolor intracellular analysis, especially when the experimental question depends on how cells are physically organized rather than only on marker presence.
- Key Risks and Design Considerations: The main design risk in cytoskeleton staining is overinterpreting brightness or apparent structural change without confirming that the staining pattern is preserved consistently across samples. Fixation quality, permeabilization strength, cell density, and imaging settings can all alter how filament organization appears. For robust interpretation, cytoskeleton staining should be evaluated in relation to cell morphology, processing consistency, and whether the assay is meant to support descriptive imaging, comparative structural analysis, or integration into a broader multicolor workflow.
Exploring stains for mitochondria, nucleus, membrane, and other organelles?
We can help you compare organelle-specific staining strategies based on target structure, live-cell compatibility, imaging duration, and multiplexing needs.
How to Choose Fluorescent Dyes for Different Organelle Cell Staining?
Choosing fluorescent dyes for organelle-specific cell staining should begin with the biological role of the target organelle rather than with color preference alone. Different organelles require different staining logic because their accessibility, membrane properties, internal environment, and compatibility with live-cell or fixed-cell workflows are not the same. In practice, researchers in pharma, biotech, and drug discovery settings often need to evaluate several factors at once, including whether the dye is intended for structural localization or function-sensitive readout, whether the workflow is live or endpoint-based, whether the signal must survive additional processing, and whether the stain will be used alone or as part of a multicolor panel. The table below provides a practical selection framework that links major organelle targets with their typical staining logic, workflow considerations, and dye-selection priorities.
Table 1. Practical framework for choosing fluorescent dyes for different organelle staining targets.
| Organelle Target | Main Staining Goal | Typical Staining Logic | Key Dye Selection Priorities | Workflow Considerations |
|---|---|---|---|---|
| Nucleus | Cell counting, segmentation, spatial reference, and nuclear morphology analysis. | Usually based on strong interaction with DNA-rich structures. | Clear nuclear contrast, appropriate permeability, clean spectral placement, and compatibility with image analysis. | Choose according to whether the experiment requires live-cell labeling, fixed-cell endpoint analysis, or a stable multicolor reference channel. |
| Mitochondria | Organelle localization, morphology comparison, and in some workflows functional state-related interpretation. | Can be localization-oriented or partly dependent on mitochondrial physiological properties. | Decide whether the dye should report structure only or also respond to organelle state; prioritize signal clarity and workflow-matched retention. | Especially important to distinguish live-cell imaging needs from post-fix workflows, because signal meaning may change after processing. |
| Plasma Membrane | Boundary definition, morphology analysis, segmentation support, and surface-oriented imaging. | Usually based on membrane association or membrane-partitioning behavior. | Strong boundary definition, low diffuse intracellular spread, suitable brightness, and compatibility with adjacent channels. | Consider whether the membrane stain is needed for live-cell tracking, endpoint morphology, or multicolor structural reference. |
| Lysosomes | Acidic compartment visualization, trafficking studies, and organelle interaction analysis. | Often depends on acidic compartment accumulation or enrichment. | Select dyes with clear punctate signal, appropriate sensitivity to compartment conditions, and compatibility with the biological question. | Signal can be highly sensitive to sample condition and handling, so validation under real workflow conditions is important. |
| Endoplasmic Reticulum | Intracellular network visualization and internal architecture analysis. | Designed to reveal extended reticular membrane organization rather than compact organelle signal. | Prioritize coherent network-like pattern, manageable background, and compatibility with broader intracellular panels. | Evaluate whether the stain performs better in live imaging or fixed endpoint analysis, especially when network continuity matters. |
| Golgi Apparatus | Spatial organization analysis and compact intracellular reference staining. | Typically relies on localized intracellular patterning associated with Golgi distribution. | Choose dyes that provide compact, well-resolved localization rather than broad cytoplasmic fluorescence. | Processing effects can alter apparent localization, so stability and reproducibility across conditions should be checked carefully. |
| Lipid-Associated Compartments | Visualization of membrane-rich domains, lipid-related structures, and lipid-environment changes. | Often based on lipid interaction, membrane partitioning, or environment-sensitive fluorescence behavior. | Clarify whether the need is broad membrane-associated contrast or more discrete lipid-compartment interpretation. | These stains are often highly context-dependent, so sample type, treatment condition, and live-versus-fixed workflow all matter strongly. |
| Multiorganelle Panels | Simultaneous comparison of several intracellular structures in one experiment. | Combines structural and sometimes function-sensitive channels across multiple organelles. | Prioritize spectral separation, brightness balance, panel compatibility, and control-friendly channel assignment. | Select dyes not only by organelle target, but also by how each channel performs within the full multicolor workflow. |
How to Combine Organelle Stains in Multicolor Experiments?
Many research workflows do not rely on a single organelle channel. Instead, they combine nucleus, membrane, mitochondria, lysosome, ER, Golgi, or lipid-associated channels in one experiment to build a more complete intracellular map. This can substantially improve interpretive power in microscopy, phenotypic screening, and mechanism-oriented imaging studies, but only when the panel is designed as an integrated system. If fluorophores are added without considering signal hierarchy, channel spacing, and the biological role of each stain, the result may be visually complex but analytically weak.
- How to Assign Organelle Stains Across Different Channels: Channel assignment should begin with analytical priority rather than with color preference alone. Structural anchor channels such as the nucleus or plasma membrane usually need to remain clear and stable, but they do not always need the cleanest or brightest spectral positions if those channels are required for weaker or more biologically sensitive organelle readouts. In practice, channel selection should reflect which targets are most critical to the biological question, which organelles are expected to produce weaker signal, and which channels must remain the most interpretable after acquisition and analysis.
- Avoiding Spectral Overlap and False Co-Localization: One of the most important technical requirements in multiorganelle staining is preventing spectral overlap from being misread as biological co-localization. This is especially important when the experiment is designed to study organelle interaction, redistribution, or compartment proximity. Fluorophores should therefore be selected with enough spectral separation that overlapping signals do not produce artificial spatial association. Single-stain controls and careful channel validation are essential because even visually attractive multicolor images can contain bleed-through that undermines the biological conclusion.
- Balancing Structural Reference Dyes and Functional Organelle Probes: Not every organelle channel plays the same role in a multicolor panel. Some channels are primarily structural references, such as nuclear or membrane stains, while others may be more sensitive to compartment state, such as certain mitochondrial or lysosome-related probes. These channels should not be designed or interpreted as though they are equivalent. A robust panel preserves both structural clarity and functional meaning by assigning channels according to what each stain is actually expected to contribute to the final interpretation.
- Controls Needed for Interpreting Multiorganelle Staining Panels: Multiorganelle panels require stronger validation than single-channel experiments because complexity increases the chance of false interpretation. Unstained controls, single-stain controls, and stable acquisition settings are critical for distinguishing real compartment-specific signal from background, spillover, or processing-related changes. In practice, the more channels added to the experiment, the more important it becomes to treat control design as part of panel architecture rather than as an optional confirmation step.
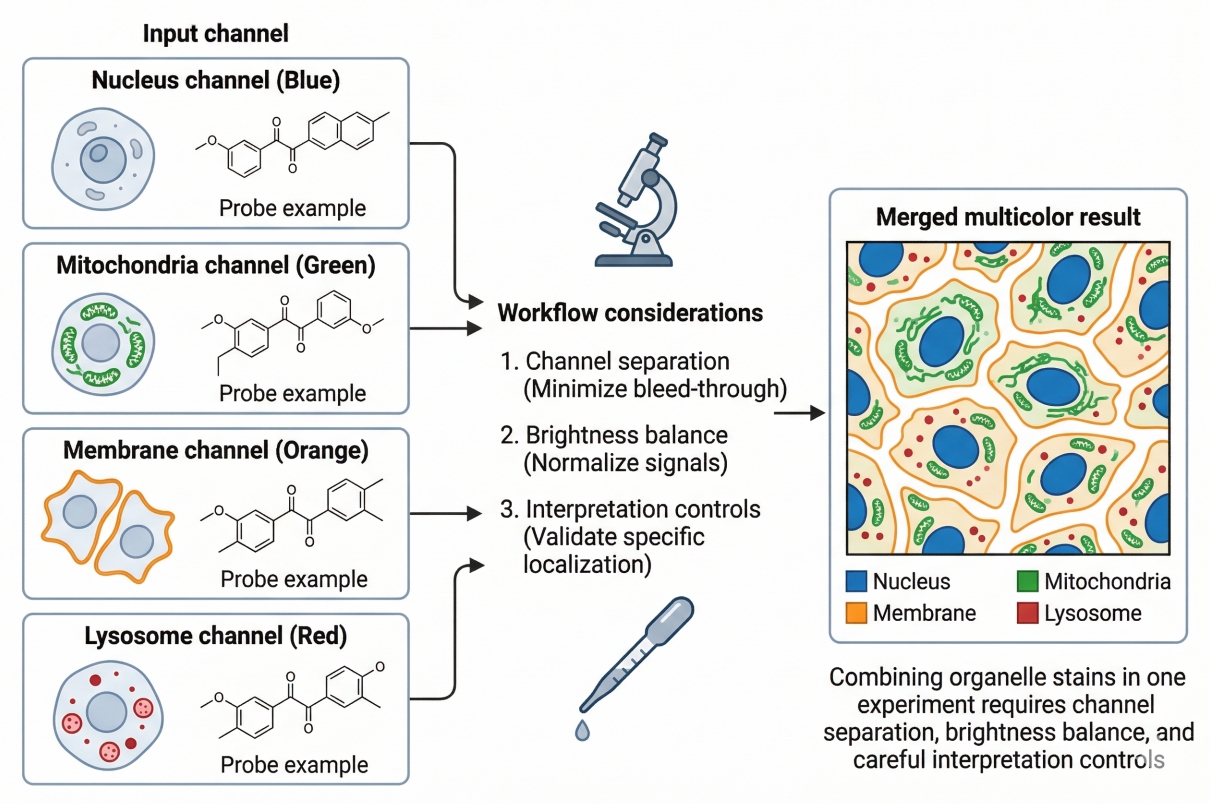 Fig. 2. Combining organelle stains in one experiment requires channel separation, brightness balance, and careful interpretation controls.
Fig. 2. Combining organelle stains in one experiment requires channel separation, brightness balance, and careful interpretation controls.
Common Pitfalls in Organelle Cell Staining and How to Avoid Misinterpretation
Organelle-specific staining provides rich intracellular information, but it is also highly vulnerable to overinterpretation when signal behavior is treated too simply. The most common problems do not usually arise because no fluorescence is visible. They arise because the fluorescence looks plausible while still failing to support a reliable biological conclusion. In organelle-focused imaging, avoiding misinterpretation is often more important than maximizing brightness, because many apparently strong signals become misleading when probe mechanism, workflow design, and processing effects are not considered together.
- Mistaking Dye Accumulation for True Organelle Specificity: A bright or localized signal does not automatically prove that a probe is truly organelle-specific under the chosen experimental conditions. Some dyes accumulate strongly because of physicochemical preference, membrane association, or sample condition rather than because they are uniquely restricted to one intracellular compartment. This can create images that appear highly convincing while still lacking the specificity required for confident interpretation. The safest approach is to evaluate staining logic, expected localization pattern, and control performance together rather than using brightness alone as evidence of specificity.
- Misreading Functional Dyes as Purely Structural Markers: Many organelle-related probes are not purely anatomical labels. Their signal may depend partly on compartment state, such as acidity, membrane potential, or lipid environment. If these probes are interpreted as though they only show where an organelle is, changes in fluorescence may be overread as structural redistribution or altered abundance when they actually reflect a shift in organelle condition. This is especially important in treatment-response and mechanistic studies where fluorescence changes are often central to interpretation.
- Signal Changes Caused by Fixation, Washing, or Sample Stress: Organelle-associated fluorescence can change substantially during fixation, permeabilization, aggressive washing, or because the sample itself has been stressed before imaging. These workflow-related changes may alter localization, retention, intensity, or background in ways that resemble biology. A probe that performs well in live cells may lose interpretive value after fixation, and a compartment-sensitive signal may shift because handling changed the sample state rather than because the biological target changed. For this reason, organelle staining results should always be interpreted in the context of the exact workflow used to generate them.
- A Practical Framework for Choosing the Right Organelle Stain: The most reliable way to avoid misinterpretation is to choose organelle stains through a structured framework rather than by color or familiarity alone. Researchers should define which compartment must be visualized, whether the readout is primarily structural or partly functional, whether the sample will remain live or move into fixed processing, and whether the stain must be integrated into a multicolor panel. Once those questions are answered, stain selection becomes much more rational and much less vulnerable to technically impressive but biologically weak results.
Organelle-Specific Cell Staining Solutions at BOC Sciences
BOC Sciences provides comprehensive organelle-specific cell staining support to meet diverse intracellular imaging and fluorescence analysis needs in pharma, biotech, drug discovery, and CDMO research workflows. Whether the goal is nucleus visualization, mitochondrial tracking, membrane definition, lysosome analysis, ER or Golgi localization, or lipid-associated compartment imaging, we offer relevant fluorescent product resources and workflow-oriented support to help researchers build more reliable staining strategies. Our support is designed to improve organelle-level signal specificity, multicolor compatibility, and experimental reproducibility across both live-cell and fixed-cell applications.
Organelle-Targeted Fluorescent Products for Intracellular Imaging
- Offers organelle-focused product categories including nuclear fluorescent probes, mitochondrial fluorescent probes, lysosomal fluorescent probes, endoplasmic reticulum fluorescent probes, Golgi fluorescent probes, cell membrane fluorescent probes, and lipid fluorescent probes.
- Provides broader staining-related product resources such as fluorescent dyes, fluorescent probes, cell and organelle stains, and specific fluorescent stains for projects requiring wider fluorophore screening or cross-category comparison.
- Covers visible to longer-wavelength fluorescence options that can support single-color and multicolor organelle imaging workflows across different microscopy and fluorescence detection platforms.
- Supports projects ranging from single-organelle visualization to multiorganelle imaging panels for intracellular localization and comparative phenotype analysis.
Workflow-Matched Dye Selection for Live-Cell and Fixed-Cell Applications
- Supports selection of fluorescent products according to whether the experiment is designed for live-cell staining, fixed-cell imaging, or workflows that combine early live observation with later endpoint analysis.
- Helps align stain choice with organelle-specific staining logic, including structure-oriented labeling, membrane-associated localization, acidic compartment staining, and environment-sensitive organelle readouts.
- Provides product direction for assays that require strong signal retention, reduced nonspecific background, and compatibility with washing, fixation, or multistep intracellular imaging workflows.
- Supports selection strategies that take into account imaging duration, fluorescence stability, and practical assay conditions in dynamic or endpoint organelle analysis.
Multicolor Organelle Imaging and Fluorophore Planning Support
- Provides fluorophore resources relevant to multicolor organelle staining workflows where nuclei, membranes, mitochondria, lysosomes, ER, Golgi, or lipid-associated structures need to be analyzed in combination.
- Supports stain selection with attention to spectral spacing, channel compatibility, and fluorophore balance for microscopy-based multicolor imaging experiments.
- Helps research teams identify product directions that are more suitable for structural reference channels versus organelle probes with stronger sensitivity to intracellular state or compartment environment.
- Supports broader panel-planning needs in workflows where intracellular co-localization, organelle interaction, or multichannel phenotype comparison is required.
Expanded Fluorescence Resources for Customized Research Needs
- Offers related fluorescence-enabling product categories such as NHS esters, hydrazides, azides, alkynes, and click chemistry reagents for research teams developing more customized fluorescent reagents or labeling workflows.
- Provides access to related fluorescence support resources such as fluorescent protein services, fluorescent nanoparticle services, and fluorescent bead services for projects extending beyond standard organelle stain selection.
- Supports integration of organelle staining with broader research areas including cell imaging, fluorescence imaging, molecular imaging, and high-throughput screening.
- Enables research teams to build more scalable fluorescence workflows for intracellular imaging, phenotypic profiling, assay development, and multiorganelle analysis.
Do You Need A Consultation?
BOC Sciences integrates cutting-edge fluorescence technologies to accelerate your research, driving next-generation solutions for drug discovery and diagnostics.
Transform Your Studies with Cutting-Edge Fluorescent Products
| Catalog | Name | CAS | Inquiry |
|---|---|---|---|
| A16-0076 | Astrophloxine | 14696-39-0 | Bulk Inquiry |
| A16-0012 | Lucigenin | 2315-97-1 | Bulk Inquiry |
| A16-0028 | Monodansylcadaverine | 10121-91-2 | Bulk Inquiry |
| A16-0049 | C12 NBD Galactosylceramide | 474942-98-8 | Bulk Inquiry |
| A16-0074 | Biotin DHPE | 136235-58-0 | Bulk Inquiry |
| A16-0153 | NBD cholesterol | 78949-95-8 | Bulk Inquiry |
| A16-0073 | NBD Dihexadecylamine | 117056-66-3 | Bulk Inquiry |
| A16-0183 | NBD-C6-HPC | 81005-34-7 | Bulk Inquiry |
| A16-0206 | C-Laurdan | 959839-06-6 | Bulk Inquiry |
| A16-0194 | LysoTracker Red DND-99 | 231946-72-8 | Bulk Inquiry |
High-Performance Fluorescent Tools for Your Research
- Nuclear Fluorescent Probes DNA-binding dyes for nucleus visualization.
- Endoplasmic Reticulum Fluorescent Probes ER-targeted dyes for organelle structure analysis.
- Lipid Fluorescent Probes Dyes for lipid droplets and membranes.
- pH Indicators Fluorescent sensors for intracellular pH monitoring.
- Mitochondrial Fluorescent Probes Targeted dyes for mitochondrial structure and function.
- Cytoskeleton Fluorescent Probes Probes for actin and microtubule visualization.
- Metal Fluorescent Probes Selective sensors for intracellular metal ions.
- Cell membrane Fluorescent Probes Surface-labeling dyes for membrane dynamics studies.
Explore More Topics
Online Inquiry