Thiazole Orange (TO) Fluorescent Staining Protocols: Nucleic Acid Fluorescent Dye
Thiazole Orange (TO) is a highly sensitive fluorescent dye that binds strongly to nucleic acids, producing a pronounced increase in fluorescence upon interaction with DNA or RNA. Its ability to label both single- and double-stranded nucleic acids makes TO an ideal tool for detecting cellular nucleic acid content, performing flow cytometry analyses, and constructing nucleic acid probe systems. With its high brightness and low background, TO enables real-time imaging and accurate nucleic acid detection across a variety of experimental applications.
Molecular Structure of Thiazole Orange
Thiazole Orange belongs to the class of asymmetric cyanine dyes, consisting of two heteroaromatic rings (quinoline and benzothiazole) connected by a carbon–carbon double bond. The nitrogen atom on the quinoline ring carries a positive charge, conferring water solubility and enabling electrostatic interaction with the negatively charged nucleic acid backbone. The dye features an extended planar conjugated system, allowing intercalation between DNA base pairs. TO is typically supplied as the hydrochloride or p-toluenesulfonate salt, with a molecular weight of approximately 476.6. Its maximum absorption occurs around 509 nm, with an emission peak at 532 nm, corresponding to green fluorescence (excitation/emission spectra similar to fluorescein).
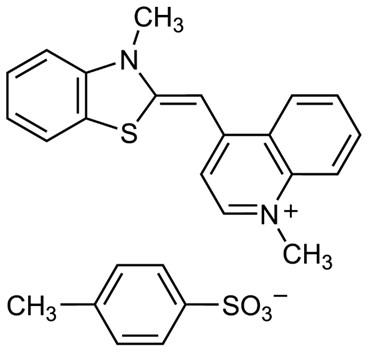 Fig. 1. Molecular structure of thiazole orange nucleic acid fluorescent dye (BOC Sciences Authorized).
Fig. 1. Molecular structure of thiazole orange nucleic acid fluorescent dye (BOC Sciences Authorized).
Development History of Thiazole Orange
In the mid-1980s, researchers designed and synthesized Thiazole Orange based on the structure of Thioflavin T to enable fluorescent flow cytometric counting of reticulocytes. In a pioneering study by Lee et al. in 1986, it was discovered that TO can be excited at 488 nm and provides superior staining compared to traditional dyes such as acridine orange. More importantly, the fluorescence intensity of TO increases dramatically upon binding to nucleic acids. Free TO exhibits minimal fluorescence, whereas binding to cellular RNA/DNA can enhance fluorescence by over 3000-fold, resulting in an exceptionally high signal-to-noise ratio. This pronounced "light-up" property quickly drew attention, demonstrating TO's ultra-sensitive detection of double-stranded nucleic acids.
By the 1990s, TO became widely used in nucleic acid experiments, gradually replacing some conventional dyes. For instance, in agarose gel electrophoresis, TO serves as a general nucleic acid stain, sensitively visualizing DNA/RNA bands. In blood and cultured cell analyses, TO was employed for total nucleic acid staining, such as detecting reticulocytes or lymphocyte proliferation via flow cytometry. Compared to ethidium bromide (EB), which requires UV excitation and exhibits higher toxicity, TO's visible-light excitation and lower mutagenicity (Ames test indicates ~1/3–1/4 the mutagenic potential of EB) offered significant advantages, contributing to its widespread adoption in research and teaching laboratories. Researchers also developed a series of TO derivatives and improved versions. For example, in the 1990s, Molecular Probes introduced covalently linked TO dimers TOTO-1 and YOYO-1 to enhance binding to long DNA molecules. Additionally, substituent modifications on the TO structure led to the SYBR Green family of commercial dyes for PCR product detection.
Entering the 21st century, TO applications expanded from non-specific dyes to sequence-specific probes. Covalent attachment of TO to oligonucleotides enabled novel probes. For example, in 2000, Kubista et al. designed TO-labeled peptide nucleic acid (PNA) "Light-up probes" that enhanced fluorescence up to 50-fold upon binding target DNA for homogeneous real-time PCR detection. Subsequently, the Seitz group developed FIT probes by embedding TO into conventional oligonucleotides, and the Okamoto group constructed ECHO probes utilizing TO dimer-induced Förster resonance energy transfer (FRET) for imaging specific RNA in living cells. These pioneering studies illustrate TO's evolution from a general dye to a functionalized probe. Today, TO and its derivatives remain crucial in nucleic acid analysis and fluorescence sensing. Due to its extremely low background fluorescence and robust signal activation, TO is widely used in new sensing systems for detecting various biomacromolecules and metal ions.
Mechanism of Action of Thiazole Orange Staining
The fluorescence intensity of TO is highly dependent on its molecular conformation. When free in solution, the quinoline and benzothiazole rings can freely rotate around the central double bond, placing the dye in a non-planar twisted conformation that disrupts conjugation. Excitation energy is predominantly dissipated non-radiatively, resulting in very weak background fluorescence (quantum yield<0.001, nearly undetectable). Upon binding to DNA, the situation changes dramatically. TO intercalates between base pairs of double-stranded DNA, where base stacking and steric constraints force the molecule into a planar conjugated conformation. In this configuration, the π orbitals of the aromatic rings are fully conjugated, allowing efficient radiative energy release and bright fluorescence emission. Reports indicate that TO's quantum yield increases to ~0.4 upon binding double-stranded DNA, representing a several-thousand-fold enhancement relative to the free state.
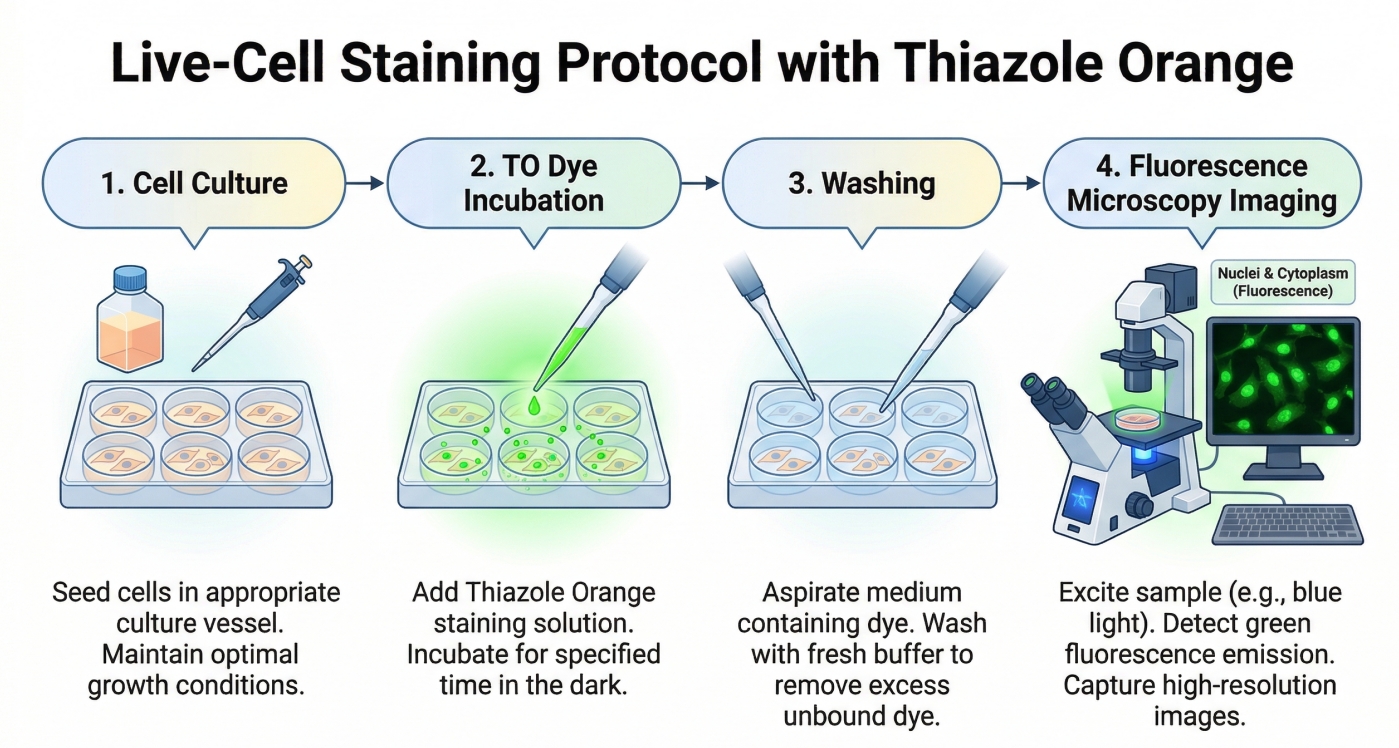 Fig. 2. A diagram illustrating the four-step Thiazole Orange staining protocol (BOC Sciences Authorized).
Fig. 2. A diagram illustrating the four-step Thiazole Orange staining protocol (BOC Sciences Authorized).
TO binds nucleic acids primarily via two modes: intercalation and groove binding. In double-stranded DNA, intercalation predominates, with the planar TO molecule inserting between adjacent base pairs and stacking perpendicularly to the bases, forming a stable complex. Intercalation anchors TO away from solvent quenching, producing strong green fluorescence. The positive charge of TO further strengthens binding via electrostatic interaction with the DNA phosphate backbone. In certain conditions, such as irregular nucleic acid structures or high dye/base ratios, TO may partially bind along the DNA groove, resulting in weaker fluorescence enhancement compared to intercalation. Overall, TO exhibits high selectivity for double-stranded nucleic acids. Regular double-helical structures provide optimal intercalation sites, yielding marked fluorescence enhancement, whereas single-stranded nucleic acids bind TO weakly, producing minimal fluorescence increase. Consequently, TO is commonly used to detect double-stranded nucleic acids and distinguish between single- and double-stranded states. Additionally, TO's excitation and emission wavelengths are in the visible range (typical excitation ≈510 nm, emission ≈530 nm), allowing excitation with a 488 nm laser or blue LED and emission of green fluorescence. Compared to dyes requiring UV excitation, visible-light excitation reduces sample autofluorescence and minimizes photodamage to cells and DNA, representing a major advantage in both mechanism and application.
Staining Procedure of Thiazole Orange
Experimental Steps
Fluorescence Microscopy Staining (TO Nucleic Acid Staining):
- Prepare working solution: Dilute TO in PBS to 0.1–1 µM (from a 10 mM DMSO stock solution if needed).
- Sample incubation:
- Immerse live or fixed cells in the TO working solution and incubate at room temperature for 5–30 minutes (adjust according to sample type and dye concentration).
- For live cell staining, use lower concentrations and shorter incubation times to minimize impact on cell function; for fixed cells, longer incubation may be used to achieve uniform staining.
- Washing: Rinse with fresh buffer to remove unbound dye and reduce background.
- Fluorescence observation:
- Stained nuclei emit green fluorescence in the FITC/GFP channel.
- Can be combined with other fluorescent dyes (e.g., DAPI for blue nuclear staining or red mitochondrial dyes), taking care to avoid spectral overlap interference.
Nucleic Acid Gel Staining:
a. Gel-embedded (Pre-staining)
- Prepare gel: Add TO to agarose gel solution to a final concentration of approximately 1.0–1.3 µg/mL (2–3 µM).
- Electrophoresis: Perform standard agarose gel electrophoresis; DNA bands can be directly visualized post-run without additional processing.
b. Post-staining
- Gel incubation: Immerse the gel in TE buffer containing ~1 µg/mL TO and gently shake for 20–30 minutes.
- Washing: Rinse with water to remove free dye and reduce background.
- Fluorescence observation:
- Visualize bright green fluorescence under UV or blue light (recommended excitation 470–490 nm with a green filter).
- Avoid UV-induced DNA damage, facilitating downstream cloning experiments.
- Safety note: TO aqueous solutions are relatively stable and less mutagenic, but gels and waste solutions should still be handled according to standard laboratory chemical safety guidelines.
Real-time Fluorescent PCR Detection (qPCR):
- Prepare TO solution: Make a high-concentration stock (e.g., 10 mM in DMSO) and dilute to a final concentration of ~0.2–1 µM in the PCR reaction.
- Real-time detection: During each extension cycle, newly synthesized dsDNA binds TO, increasing fluorescence for real-time monitoring of product accumulation.
- Precautions:
- Avoid excessively high TO concentrations, which may inhibit DNA polymerase or affect amplification efficiency.
- Intercalation of TO may slightly increase DNA melting temperature.
- Special applications:
- TO can be covalently attached to primers or probes (e.g., via NHS ester activation to amino-modified oligonucleotides) to improve specificity.
- Upon hybridization with the target sequence, TO intercalates into the double-stranded region, generating specific fluorescence for detection of particular genes or mutations.
Combined Staining and Multiplex Detection:
a. TO + PI Dual Staining (Live/Dead Cell Assay)
- Preparation: Dilute TO and PI each to ~1 µM and add to the cell suspension.
- Incubation: Protect from light and incubate for 5–10 minutes.
- Observation/Analysis:
- Live cells exhibit green fluorescence (TO); dead cells show red fluorescence (PI).
- Flow cytometry can quantify cell populations based on green/red fluorescence.
b. Multiplex Fluorescence Staining
- Nucleic acid multiplexing: Stain nuclei with TO (green) while simultaneously using DAPI or Hoechst 33342 (blue) for other nucleic acid components.
- Protein immunofluorescence: Use TO nucleic acid staining as background while labeling proteins with red fluorescent dyes to visualize cellular structures.
- Precautions:
- Avoid competition or energy transfer interference between nucleic acid dyes (e.g., TO and other iodinated dyes).
- If GFP or other green fluorescent proteins are present, avoid TO to prevent signal confusion.
- Optimize dye concentrations and staining order in multiplex experiments through control tests.
Usage Considerations
- Photostability and Photobleaching: TO is not extremely light-sensitive, but prolonged exposure to strong light can lead to fluorescence decay and dye degradation. Minimize exposure to daylight or excitation light. Use anti-fade mounting media during microscopy and shorten exposure times to reduce photobleaching. Store TO in the dark at low temperature (e.g., 4°C) to prolong shelf life.
- Background Staining and Non-specific Binding: Although free TO has weak fluorescence, background signals may arise in practice. In cell staining, residual unwashed TO may mildly stain membranes, extracellular matrix, or culture medium components, generating uniform background. TO's positive charge can also weakly interact with other negatively charged molecules (e.g., certain proteins, polysaccharides, or cell debris), causing non-specific fluorescence. Appropriate post-staining washing and control experiments are recommended to confirm signal specificity. For quantitative studies (e.g., flow cytometry), subtract the background of unstained controls. For thick tissue sections, extend washing to remove unpenetrated dye and prevent surface-background fluorescence.
- Dye Aggregation and Self-quenching: At high concentrations, TO molecules may undergo π-stacking aggregation, leading to fluorescence quenching or spectral changes. This is more likely in the absence of nucleic acids. For instance, overly concentrated stock solutions may exhibit slight background fluorescence or turbidity. To avoid this, prepare working solutions according to product instructions, optionally dissolving TO in DMSO first and then gradually diluting into aqueous buffer. Avoid leaving TO in high-concentration nucleic acid solutions for prolonged periods, as excess TO may self-aggregate once binding sites are saturated. For higher staining intensity, consider TO derivatives with steric hindrance groups to reduce dye-dye interactions.
- Biocompatibility and Competitive Binding: As a nucleic acid intercalating dye, TO has some cytotoxicity and potential genotoxicity in live cells. Although less mutagenic than ethidium bromide, TO can interfere with DNA replication and RNA transcription; prolonged or high-dose exposure may affect cell proliferation. In live-cell experiments, minimize concentration and staining time, using TO only for end-point observation. For dynamic live-cell imaging, assess TO's effect on cell physiology or select lower-toxicity alternatives. Additionally, in systems containing other nucleic acid-binding reagents (e.g., histones, DNA-binding drugs, or other dyes), TO binding may be competitively inhibited, reducing staining efficiency or signal intensity. Conversely, TO may affect the function of other molecules. In experiments combining multiple dyes or antibiotics, evaluate potential competition for nucleic acid binding. To ensure accuracy, test each condition separately and adjust TO concentration as needed. In multiplex experiments, prevent TO fluorescence from causing "bleed-through" in other channels; spectral compensation or post-processing may be applied if necessary.
Usage Limitations
- Live-cell and real-time imaging limitations: TO can freely penetrate cell membranes without requiring fixation, making it suitable for live-cell staining. However, in live cells, TO cannot target specific nucleic acid sequences; it stains all DNA and RNA simultaneously, including nuclear DNA, mitochondrial DNA, and ribosomal RNA. Therefore, fluorescence observed in live-cell microscopy reflects the total nucleic acid content, making it difficult to resolve specific genes or subcellular regions. Additionally, TO's intercalation and stabilization of double helices may disrupt normal nucleic acid metabolism in live cells, and prolonged staining may inhibit cell proliferation. For these reasons, TO is better suited for transient staining and end-point analysis rather than long-term live-cell tracking of nucleic acid dynamics.
- Poor sequence and structure selectivity: TO itself lacks sequence specificity and binds to any source of double-stranded DNA or RNA, enhancing fluorescence non-selectively. Experiments requiring identification of specific sequences (e.g., detecting a viral gene or single-base mutations) cannot rely on direct TO staining; sequence-specific probes (such as TO-labeled oligonucleotide probes) are needed to achieve selectivity. Similarly, TO does not strongly discriminate between different higher-order structures (B-DNA, A-RNA, G-quadruplexes, triplex DNA, etc.). Any nucleic acid with stacked bases or double-stranded regions may bind TO and fluoresce. This non-selective nature limits TO's use in fine structural analysis and often requires combination with sequence-specific techniques for selective detection.
- Non-specific background and intracellular localization: TO preferentially binds double-stranded regions, which can lead to non-target fluorescence in complex biological samples. In tissue sections or cell samples, TO stains not only genomic DNA but also accumulates in nucleoli (rich in rRNA duplexes) and mitochondria (circular double-stranded DNA), producing strong fluorescence that may obscure other details. When the region of interest is not nucleoli or mitochondria, this high background can interfere with analysis. In microbial applications, TO labels all microbial DNA equally, preventing species-specific differentiation. These considerations indicate that TO is more suitable as a general-purpose stain, while experiments requiring high specificity should use modified TO derivatives with targeting groups.
- Tissue staining and imaging depth: TO staining is most effective in isolated cells or thin samples, with limitations in thick tissues or in vivo applications. TO molecules, though moderately sized, require sufficient diffusion to penetrate hundreds of micrometers into tissue, often necessitating longer incubation or tissue clearing/permeabilization procedures. Even then, the central regions of thick tissue blocks may not receive adequate dye. Moreover, TO's excitation wavelength (~510 nm) in the visible green range limits tissue penetration and is prone to scattering and autofluorescence, making deep-tissue signal acquisition difficult. Therefore, TO is typically used for cultured cells or thin tissue sections rather than whole organs or tumors. For deep-tissue imaging, near-infrared dyes or probes are generally preferred. Overall, TO's spatial application is limited, favoring in vitro and microscopy-level analysis over non-invasive in vivo labeling.
Applications of Thiazole Orange Staining
Nucleic Acid Fluorescence Staining
TO is commonly used as a general DNA/RNA fluorescent dye. For example, adding TO to agarose gels allows direct visualization of nucleic acid bands after electrophoresis. It is also used for staining nucleic acids in cells or tissue sections under fluorescence microscopy. TO can penetrate cell membranes and bind both DNA (nucleus) and RNA (cytoplasm, e.g., ribosomal RNA), producing green fluorescence for nuclear or total nucleic acid staining. In flow cytometry, TO is used for reticulocyte counting (binding residual RNA) and cell cycle analysis (combined with RNase to distinguish DNA vs RNA signals).
Real-time PCR and Nucleic Acid Detection
TO is an important fluorescent component in real-time qPCR and molecular probe technologies. It can be added directly to PCR amplification reactions, binding newly synthesized dsDNA to produce fluorescence similar to SYBR Green, enabling real-time monitoring of product accumulation. Due to its strong affinity for double-stranded nucleic acids and low background, TO provides a high signal-to-noise ratio. Additionally, TO can be covalently attached to oligonucleotide probes as a reporter molecule. For example, in molecular beacon hairpin probes, TO is positioned near a quencher and emits strong fluorescence only when the probe hybridizes with the target sequence. In Light-up and FIT probes, TO is linked directly to the probe; free in solution, fluorescence is quenched, but upon target hybridization, TO intercalates into the double-stranded region, generating a detectable signal. These applications make TO a versatile reporter for in vitro nucleic acid detection (real-time PCR, isothermal amplification) and in vivo imaging (e.g., mRNA tracking).
Cell Membrane Permeability and Live-Cell Analysis
With its small molecular size and positive charge, TO can penetrate live-cell membranes, making it suitable for viability and membrane-permeability studies. A classical application is TO/PI dual staining: TO stains total nucleic acids in all cells, while PI only enters membrane-compromised dead cells. Under flow cytometry or fluorescence microscopy, live cells show green fluorescence (TO only), and dead cells appear yellow/red (TO + PI). This method is widely used to assess cell viability, membrane integrity under drug treatment, and, in microbiology, to count total microbial cells and distinguish live/dead populations using complementary dyes.
FRET Sensors and Other Probes
TO is also employed in constructing fluorescent sensors and FRET probe systems. It can act as an energy donor or acceptor in FRET pairs; for instance, TO-labeled oligonucleotide probes can transfer excitation energy to another dye-labeled probe, reporting nucleic acid folding or binding events. Additionally, TO's "light-up" property is used in small-molecule sensors. TO-derivative probes detect metal ions or enzymatic activity by switching from free-rotating to planar fluorescent states upon target binding. TO also interacts specifically with special nucleic acid structures such as triplex DNA or G-quadruplexes, stabilizing these structures and enhancing fluorescence for structural recognition and sensor development.
Fluorescent Dyes Recommended for Your Research Project
| Catalog | Name | CAS | Inquiry |
|---|---|---|---|
| A19-0006 | Thiazole Orange | 107091-89-4 | Bulk Inquiry |
| A19-0101 | Propidium Iodide | 25535-16-4 | Bulk Inquiry |
| A19-0012 | YOYO-3 | 156312-20-8 | Bulk Inquiry |
| A19-0014 | TO-PRO3 iodide | 157199-63-8 | Bulk Inquiry |
| F02-0132 | Thiazole Orange iodide | 24147-36-2 | Bulk Inquiry |
| F02-0130 | SYBR Gold | 213019-16-0 | Bulk Inquiry |
| F02-0129 | SYBR Safe | 1030826-36-8 | Bulk Inquiry |
| A19-0087 | DusQ 2 phosphoramidite | 374591-98-7 | Bulk Inquiry |
| A19-0096 | PICO green | 178918-98-4 | Bulk Inquiry |
| A19-0094 | R6G phosphoramidite, 6-isomer | 1355330-47-0 | Bulk Inquiry |
Corporate Profile
BOC Sciences is a trusted provider of advanced fluorescent chemicals and staining reagents, supporting a wide range of applications in life sciences, biotechnology, and analytical research. With strong expertise in fluorescence chemistry and molecular design, the company offers high-quality nucleic acid dyes such as Thiazole Orange (TO), DAPI, Hoechst 33342, and Propidium Iodide (PI), as well as mitochondrial probes like Rhodamine 123. These reagents are widely used in cell imaging, flow cytometry, and nucleic acid detection, reflecting BOC Sciences' commitment to delivering reliable and high-performance fluorescent solutions for scientific research.
Building on its capabilities in organic synthesis and functional modification, BOC Sciences provides both standard and customized fluorescent probes with optimized spectral properties, stability, and sensitivity. The company supports the development of advanced dye systems, including SYBR Green-like nucleic acid stains, cyanine-based derivatives such as TO analogs, and specialized labeling reagents for proteins and biomolecules. Through precise structural design and rigorous quality control, BOC Sciences ensures consistent fluorescence performance, low background signal, and compatibility with diverse experimental platforms.
In addition to its extensive product portfolio, BOC Sciences offers integrated services such as custom synthesis, dye conjugation, analytical characterization, and scale-up production to meet specific research and industrial needs. Its deep technical expertise in fluorescent probe development enables tailored solutions for applications ranging from real-time PCR and molecular diagnostics to live-cell imaging and multiplex fluorescence assays. By combining scientific rigor with flexible service capabilities, BOC Sciences continues to be a reliable partner for researchers seeking high-quality fluorescent reagents and innovative solutions.
Custom Fluorescent Solutions Designed for Your Experiments
- DNA StainingPrecise fluorescent dyes for clear DNA visualization and analysis.
- Lipid StainingFluorescent solutions for effective lipid structure imaging.
- Cell StainingAdvanced staining for detailed cell morphology and analysis.
- Protein StainingHigh-quality staining for accurate protein detection and imaging.
- Bacteria ImagingFluorescent solutions to visualize bacterial structures and activity.
- Cell ImagingVisualize and analyze live or fixed cells using advanced fluorescence.
- Molecular ImagingCutting-edge fluorescent solutions for deep molecular analysis.
- Fluorescence ImagingHigh-resolution imaging solutions for detailed fluorescence studies.
- BioconjugationCustom bioconjugation services for protein, peptide, and dye linking.
- Drug DeliveryTailored fluorescent solutions for efficient drug delivery research.
- Molecular DiagnosticsFluorescent probes and markers for precise molecular diagnostics.
- Flow CytometryFluorescent dyes and reagents for enhanced flow cytometry analysis.
High-Performance Fluorescent Tools for Your Research
- Cell membrane Fluorescent Probes Surface-labeling dyes for membrane dynamics studies.
- Lipid Fluorescent Probes Dyes for lipid droplets and membranes.
- Lysosomal Fluorescent Probes Acidic organelle markers for lysosome tracking.
- Golgi Fluorescent Probes Targeted probes for Golgi apparatus visualization.
- Apoptosis Fluorescent Probes Probes detecting programmed cell death events.
- Fluorescent Probes Versatile tools for biomolecular and cellular imaging.
- Ion Fluorescent Probes Indicators for real-time ion concentration imaging.
- Metal Fluorescent Probes Selective sensors for intracellular metal ions.
Explore More Topics
Online Inquiry