Propidium Iodide (PI) Fluorescent Staining Protocols: Apoptosis Detection Dye
Propidium Iodide (PI) is a widely used red-fluorescent nucleic acid dye that plays a crucial role in cell biology research, particularly for apoptosis detection and viability assessment. Its unique property of selectively entering cells with compromised membranes makes it an ideal tool for distinguishing live from dead cells in flow cytometry, fluorescence microscopy, and microbial viability studies. Beyond viability assays, PI is frequently combined with other fluorescent probes such as Annexin V, DAPI, and Hoechst to enable multi-parameter analysis of cell death, cell cycle distribution, and DNA content, providing researchers with reliable and quantitative insights into cellular health and function.
Molecular Structure of Propidium Iodide
PI has a molecular weight of 668.4 and belongs to the phenanthridinium class of fluorescent dyes. Its rigid planar aromatic ring system allows intercalation between DNA base pairs, while the two positively charged quaternary ammonium groups form a divalent cation (commonly as a diiodide salt), providing high hydrophilicity and strong binding to negatively charged nucleic acids. At the same time, these groups prevent PI from crossing intact cell membranes. This structural feature causes a significant fluorescence enhancement upon nucleic acid binding, making PI primarily suitable for staining exposed nucleic acids in cells.
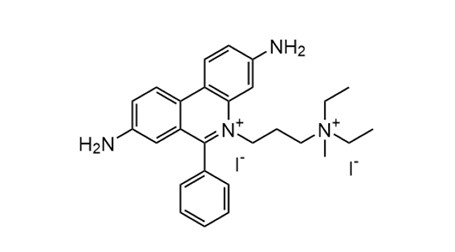 Fig. 1. Molecular structure of propidium iodide fluorescent intercalating dye (BOC Sciences Authorized).
Fig. 1. Molecular structure of propidium iodide fluorescent intercalating dye (BOC Sciences Authorized).
Historical Development of Propidium Iodide
The use of PI as a dye dates back to the mid-1970s. In 1975, Awtar Krishan first reported a flow cytometry method using PI for mammalian cell cycle analysis, considered a pioneering approach in cellular DNA quantification. Subsequently, PI quickly became a common reagent in flow cytometry for distinguishing dead cells and measuring DNA content. During the 1980s and 1990s, with the advancement of cell death studies, PI was widely applied in cell viability and cytotoxicity assays, such as evaluating antibody-mediated cytotoxic effects by differentiating live and dead cells. By the mid-1990s, methods combining PI and Annexin V staining for apoptosis detection were developed, allowing early apoptotic cells (Annexin V positive/PI negative) and late apoptotic or necrotic cells (Annexin V positive/PI positive) to be distinguished. This dual-staining approach quickly became a classic technique for assessing cell death stages.
Mechanism of Action of Propidium Iodide Staining
PI is an intercalating nucleic acid dye, with its fluorescent moiety inserting between DNA base pairs and stabilizing via electrostatic interactions with the phosphate backbone. It exhibits minimal sequence specificity and can bind double-stranded DNA or RNA similarly. The positively charged PI is attracted to negatively charged DNA and tightly intercalates into the double helix, significantly enhancing fluorescence emission. Free PI exhibits low quantum yield, whereas binding to nucleic acids increases fluorescence intensity by 20–30-fold. PI cannot penetrate intact cell membranes; therefore, it only stains cells with compromised membrane integrity, such as dead cells or cells treated with fixation/permeabilization procedures. This selectivity makes PI an ideal live/dead cell stain. Experimentally, only the nuclei of membrane-damaged cells appear red, while intact live cells remain negative.
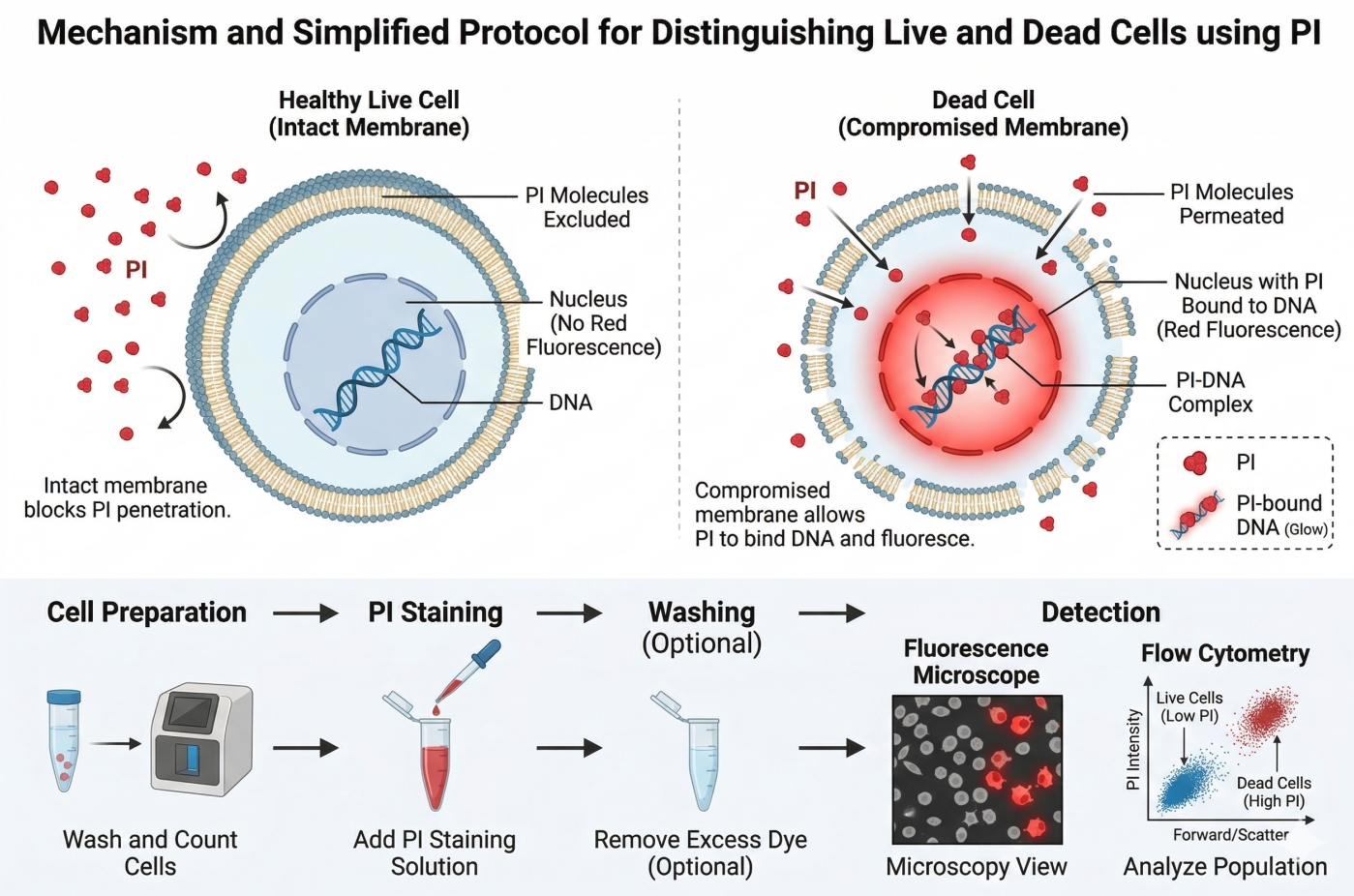 Fig. 2. Mechanism and simplified protocol of PI cell viability staining (BOC Sciences Authorized).
Fig. 2. Mechanism and simplified protocol of PI cell viability staining (BOC Sciences Authorized).
PI emits orange-red fluorescence. In aqueous solution without nucleic acids, the absorption maximum is ~493–495 nm, and the emission maximum is ~630–636 nm, with weak fluorescence. Upon intercalation into DNA, its spectrum undergoes a combination of red- and blue-shifts, with an excitation peak at ~535 nm and an emission peak at ~617 nm, producing bright orange-red fluorescence. Common excitation sources such as 488 nm lasers or green/yellow lasers (532 nm, 561 nm) effectively excite the PI-DNA complex to generate red fluorescence. In flow cytometry, PI fluorescence is typically detected in the PE-Texas Red or 610/20 nm channel. PI fluorescence intensity is proportional to the amount of bound double-stranded nucleic acids, making it suitable for quantitative DNA analysis. Since PI can also bind single- or double-stranded RNA, experiments requiring specific DNA measurement (e.g., cell cycle analysis) often involve RNase pretreatment to eliminate RNA and reduce background fluorescence.
Staining Procedure of Propidium Iodide Staining
Experimental Steps for PI Staining
Flow Cytometry Staining:
- For live cell analysis, add PI to the cell suspension at a final concentration of 5–10 μg/mL.
- Incubate on ice in the dark for 10–15 minutes.
- Keep free PI in the sample during analysis; avoid washing, as PI binding to nucleic acids is reversible, and washing may weaken the signal.
- During flow cytometry, exclude PI-positive cells to determine the percentage of live cells.
- In multicolor antibody staining, apply PI as the final step to distinguish dead cells.
- Select the appropriate fluorescence channel (e.g., 488 nm excitation, detect at 600–620 nm) to avoid spectral overlap with other dyes.
- For cell cycle analysis:
- Fix cells in 70% ethanol to permeabilize the membrane.
- Resuspend in buffer containing 50 μg/mL PI and ~0.5 μg/mL RNase A.
- Incubate at 4 °C in the dark for several hours or overnight.
- RNase A ensures DNA-only staining for accurate fluorescence.
- Acquire red fluorescence on a flow cytometer and plot DNA content histograms to calculate cell cycle phase distribution.
Fluorescence Microscopy Staining:
- For fixed cells or tissue samples, perform fixation and moderate permeabilization (e.g., 0.1% Triton X-100).
- Incubate with PI working solution (5–50 μg/mL, optimize by sample thickness) for 5–30 minutes.
- Gently wash with PBS if the sample allows; optional washing to avoid signal loss.
- Mount the sample and observe under a fluorescence microscope.
- Use PI red fluorescence as a nuclear counterstain, compatible with blue/green channel probes (e.g., FITC for target proteins).
- For live-cell confocal imaging:
- Do not use PI to stain nuclei in live cells due to toxicity.
- For monitoring cell death, add low-concentration PI to culture medium and track red nuclear fluorescence in real time.
- Control light exposure to prevent phototoxicity.
Additional Handling and Operational Tips:
- Prepare PI working solution in PBS or HBSS, optionally adding 0.1–1% BSA to reduce non-specific staining.
- Protect PI from light during storage and handling; fluorescence may degrade or produce free radicals under illumination.
- For short-term staining at room temperature, standard incubation is fine; for long-term (>30 min), store in the dark or at 4 °C to reduce background.
- In double-staining experiments:
- Be aware of dye competition between PI and other nucleic acid stains.
- For bacteria, use SYTO9 + PI as recommended by the kit.
- For eukaryotic cells, sequentially stain with Hoechst 33342/DAPI (total nuclei, blue) and PI (dead cells, red).
- Dispose of PI-containing waste properly using chlorinated disinfectants or specialized containers.
- Safety precautions: wear gloves and eye protection when handling PI powder or concentrated solutions to prevent skin or mucosal contact.
Precautions
- Phototoxicity and Safety: PI binds strongly to DNA and may generate reactive oxygen species upon light excitation, exhibiting phototoxicity and potential genotoxicity. During microscopic observation, excitation light intensity and exposure time should be minimized to prevent additional cell damage. Since PI is a mutagenic intercalator, handling the solid dye or concentrated solutions requires personal protective equipment (gloves, lab coat, and working in a fume hood) and proper disposal of residues according to biological safety protocols.
- Signal Specificity and Interference: PI staining specifically differentiates cells with intact membranes from those with compromised membranes. In multicolor experiments, spectral overlap should be avoided. PI emits broadly around >600 nm, so when used alongside PE, Texas Red, or Alexa 568/594, proper compensation or alternative dyes (e.g., 7-AAD with a more red-shifted emission) should be chosen to prevent channel crosstalk. PI can also bind intracellular nucleic acid fragments; high levels of free DNA/RNA from apoptotic cells may increase background fluorescence. Removing extracellular DNA by centrifugation or digesting with DNase can reduce nonspecific background. Studies suggest avoiding washing after PI staining because it may disrupt the dye-nucleic acid equilibrium and weaken signals. Therefore, sufficient PI should remain in the sample prior to flow cytometry acquisition to maintain stable fluorescence intensity.
- Compatibility with Other Probes: In multiplex staining, PI can generally be combined with antibodies labeling surface antigens or other organelle probes, but order and methodology are critical. Since PI requires cell entry, assays depending on intact membrane probes (e.g., calcium indicators, mitochondrial membrane potential probes) should be completed before PI staining, otherwise dead cells may interfere. For experiments involving fixation/permeabilization (cell cycle analysis, intracellular protein staining), PI is typically the last dye added. When co-staining with other nucleic acid dyes, consider competition or Förster resonance energy transfer (FRET) effects—for example, in bacterial dual staining, SYTO9 and PI compete for DNA binding, requiring adherence to standard protocols. In mammalian dual staining, complementary dye combinations (e.g., DAPI + PI, Hoechst + PI) can label total cells and dead cells simultaneously. When staining nuclei and cytoplasmic RNA concurrently, note that PI binds RNA and may obscure other RNA probe signals; DNA dyes with no RNA affinity are preferable. Overall, multiplex design should consider PI's spectral and biological properties to minimize interference.
- Sample Type Variability: Different cells and tissues may respond differently to PI. Microorganisms with thick cell walls (yeast, fungal spores) may impede PI uptake even in dead cells, requiring enzymatic or mechanical wall disruption for accurate staining. Dense tissue sections may bind some PI non-specifically; proper controls should correct for autofluorescence. In adherent cell cultures, biofilms can result in overestimation of dead cells due to extracellular DNA binding PI and covering live cells. Some cells with sublethal membrane damage may temporarily uptake PI but retain proliferation capability; weakly PI-positive cells should be interpreted with additional functional or temporal indicators.
- Cell State Dependence: PI staining depends on the physiological state of cells. Rapidly proliferating or highly metabolically active cells may show altered membrane potential or permeability, potentially causing false-positive PI staining in live cells. Conversely, early apoptotic cells may lose proliferative ability but retain membrane integrity, appearing PI-negative. PI results should be interpreted alongside morphological or additional viability markers (e.g., TUNEL staining for apoptotic DNA fragments). PI cannot distinguish cell death mechanisms; late apoptotic and necrotic cells both appear PI-positive, so additional markers such as phosphatidylserine exposure or caspase activation are needed for detailed analysis.
Limitations
- Live Cell Imaging: PI cannot stain nuclei in live cells. Intact membranes block PI entry, and high concentrations may be toxic. PI is mainly suitable for detecting dead cells or fixed/permeabilized samples. For live cell nuclear imaging (cell cycle tracking or nuclear morphology), use membrane-permeable dyes (e.g., Hoechst 33342, SiR-DNA) instead.
- DNA Specificity: PI binds nucleic acids broadly, including double-stranded RNA and ribosomal RNA. For DNA quantification (cell cycle or ploidy analysis), RNA may cause overestimation of fluorescence unless removed with RNase. Residual RNA in certain conditions (e.g., viral infection) may still affect results, requiring supplementary confirmation. DNA-specific dyes like DAPI may provide more accurate signals in these cases but require UV excitation and are not compatible with standard 488 nm flow cytometers.
- Inability to Distinguish Death Types: PI only reports membrane integrity; it cannot differentiate necrosis from late apoptosis. Early apoptotic cells with intact membranes remain PI-negative, potentially underestimating apoptosis. To identify death pathways, PI should be combined with other markers (Annexin V, mitochondrial probes).
- Sample- and Condition-Specific Constraints: Certain cell types or experimental conditions limit PI use. Large cytoplasm cells (epithelial cells, neurons) may show false positives without sufficient RNase treatment. Some cells actively expel PI (multidrug-resistant tumor cells), requiring higher concentrations or longer incubation. Microbial cell walls (Gram-positive or Gram-negative) can affect PI entry rate; optimization is needed. VBNC (viable but nonculturable) microbes may remain PI-negative despite lost proliferation, necessitating complementary methods (culture-based or metabolic dyes).
Applications of Propidium Iodide Staining
Flow Cytometry (Cell Viability Assessment)
PI is commonly used as a dead cell exclusion dye in flow cytometry. When added to a fresh cell suspension, live cells with intact membranes remain unstained, whereas dead cells with compromised membranes rapidly uptake PI, causing their nucleic acids to emit red fluorescence. Researchers can accurately determine the proportion of viable cells based on PI-positive populations. For example, in immunology and pharmacology studies, PI staining reliably distinguishes live cells from dead cells after experimental treatments.
Apoptosis Detection
For programmed cell death assays, PI is typically used in combination with Annexin V. Annexin V labels phosphatidylserine exposed on the membranes of early apoptotic cells, while PI only enters late apoptotic or necrotic cells. Dual staining allows differentiation of cells at different stages of apoptosis. The Annexin V/PI method is a classic approach for monitoring apoptosis and is widely applied in drug screening and cell biology research.
Cell Cycle Analysis
PI is also a standard probe for measuring DNA content in flow cytometry to analyze cell cycle distribution. In the classical method, cells are first fixed and permeabilized, then stained with a buffer containing PI and RNase. Because PI binding to double-stranded DNA is quantitative, each cell's fluorescence intensity is proportional to its DNA content, allowing cell populations to be distinguished from fluorescence histograms. This method was first reported by Krishan in 1975 and remains a standard tool for cell cycle research. To avoid interference from RNA staining, RNase must be included to remove RNA from the samples.
Tissue Sections and Fluorescence Microscopy
In tissue sections or fixed cell samples, PI can serve as a fluorescent nuclear counterstain. For example, in immunofluorescence of frozen or paraffin-embedded sections, DAPI or Hoechst is commonly used for nuclear staining, but PI can also be used for red fluorescent nuclear labeling. PI-stained nuclei emit red fluorescence and are often combined with probes of other wavelengths in multiplex staining experiments. For instance, Hoechst 33342 (blue, stains all nuclei) can be combined with PI (red, only marks dead cells) to assess the viability of tissues or cell samples. It is important to note that for tissue or cultured cell staining, permeabilization is required to allow PI to fully enter nuclei and stain DNA.
Microbial Viability Assessment
PI is widely used in microbiology to evaluate the viability of bacteria or fungi. For example, bacterial viability kits employ a dual-staining approach using SYTO9 and PI: SYTO9 penetrates all bacteria, staining nucleic acids green, while PI only enters membrane-compromised dead cells, staining nucleic acids red. Because PI has higher affinity for DNA, it can replace SYTO9 in dead cells, causing them to emit predominantly red fluorescence, thus distinguishing them from live bacteria (green only). Flow cytometry or fluorescence microscopy can then quantify the red/green signal ratio, reflecting the proportion of live and dead microorganisms. This approach is widely applied in environmental monitoring, water quality testing, and food fermentation studies. However, in certain cases, such as bacterial biofilms, PI staining may underestimate viable cell numbers, which should be considered when interpreting results.
Fluorescent Dyes Recommended for Your Research Project
| Catalog | Name | CAS | Inquiry |
|---|---|---|---|
| A16-0159 | HIDC iodide | 36536-22-8 | Bulk Inquiry |
| A16-0170 | Rhodamine-123 | 62669-70-9 | Bulk Inquiry |
| A19-0040 | Hoechst 33342 | 23491-52-3 | Bulk Inquiry |
| A16-0004 | Phalloidin-FITC | 915026-99-2 | Bulk Inquiry |
| A19-0101 | Propidium Iodide | 25535-16-4 | Bulk Inquiry |
| A19-0034 | DAPI dilactate | 28718-91-4 | Bulk Inquiry |
| A16-0008 | DiD perchlorate | 127274-91-3 | Bulk Inquiry |
| A16-0212 | DiO, lipophilic tracer | 28462-56-8 | Bulk Inquiry |
| A19-0017 | Hexidium Iodide | 211566-66-4 | Bulk Inquiry |
| A16-0204 | Oxazole yellow | 152068-09-2 | Bulk Inquiry |
Corporate Profile
BOC Sciences is a leading independent supplier specializing in fluorescent chemicals, dyes, and advanced staining reagents for life sciences research. With decades of experience in chemical synthesis and analytical services, we provide high-purity fluorescent probes such as Propidium Iodide (PI), DAPI, Hoechst 33342, SYTO9, and other DNA/RNA intercalating dyes. Our products are designed to ensure reliable performance in a wide range of applications, including flow cytometry, confocal microscopy, apoptosis detection, and microbial viability studies, supporting scientists in achieving accurate and reproducible experimental results.
Our expertise extends beyond standard product supply to include custom synthesis and chemical modification, allowing researchers to obtain tailored fluorophores and conjugates that meet specific experimental needs. BOC Sciences' chemists offer technical guidance on staining protocols, probe selection, and fluorescence optimization, ensuring that dyes like PI for dead cell exclusion, Hoechst for live cell nuclear staining, and SYTO9 for microbial viability assays are used effectively. With stringent quality control measures, including HPLC, NMR, and fluorescence validation, we guarantee the purity, stability, and reproducibility of all fluorescent reagents.
BOC Sciences combines scientific rigor with customer-focused service, providing comprehensive support for complex biological experiments. Our solutions enable accurate measurement of cell viability, apoptosis, DNA content, and nucleic acid localization while minimizing background interference. By integrating deep knowledge of fluorescent probe chemistry with practical laboratory experience, we help researchers optimize experimental workflows and generate robust data. This commitment to quality and innovation makes BOC Sciences a trusted partner for laboratories, academic institutions, and industrial R&D teams worldwide.
Custom Fluorescent Solutions Designed for Your Experiments
- DNA StainingPrecise fluorescent dyes for clear DNA visualization and analysis.
- Lipid StainingFluorescent solutions for effective lipid structure imaging.
- Cell StainingAdvanced staining for detailed cell morphology and analysis.
- Protein StainingHigh-quality staining for accurate protein detection and imaging.
- Bacteria ImagingFluorescent solutions to visualize bacterial structures and activity.
- Cell ImagingVisualize and analyze live or fixed cells using advanced fluorescence.
- Molecular ImagingCutting-edge fluorescent solutions for deep molecular analysis.
- Fluorescence ImagingHigh-resolution imaging solutions for detailed fluorescence studies.
- BioconjugationCustom bioconjugation services for protein, peptide, and dye linking.
- Drug DeliveryTailored fluorescent solutions for efficient drug delivery research.
- Molecular DiagnosticsFluorescent probes and markers for precise molecular diagnostics.
- Flow CytometryFluorescent dyes and reagents for enhanced flow cytometry analysis.
High-Performance Fluorescent Tools for Your Research
- Metal Fluorescent Probes Selective sensors for intracellular metal ions.
- Cell Proliferation Tracer Fluorescent Probes Long-term tracking of cell division processes.
- pH Indicators Fluorescent sensors for intracellular pH monitoring.
- Endoplasmic Reticulum Fluorescent Probes ER-targeted dyes for organelle structure analysis.
- Mitochondrial Fluorescent Probes Targeted dyes for mitochondrial structure and function.
- Cell membrane Fluorescent Probes Surface-labeling dyes for membrane dynamics studies.
- Nuclear Fluorescent Probes DNA-binding dyes for nucleus visualization.
- Ion Fluorescent Probes Indicators for real-time ion concentration imaging.
Explore More Topics
Online Inquiry